All published articles of this journal are available on ScienceDirect.
Microwave-assisted Green Synthesis and Integrated Bioinformatics Study Reveal Curcumin Analogs Dibenzylidene-cyclohexanones as Novel Potential Anti-Tuberculosis Agents
Abstract
Introduction
Tuberculosis (TB) remains a major global health challenge, further complicated by drug resistance and comorbidities. This study investigates curcumin analogues, dibenzylidene-cyclohexanones, as potential multitarget anti-TB agents.
Methods
Nine benzylidene cyclohexanone derivatives were synthesized using microwave-assisted techniques (yields: 23–81%). Their potential activities were evaluated through molecular docking against key Mycobacterium tuberculosis (Mtb) enzymes and supported by network pharmacology analysis focused on TB-related pathologies.
Results
The benzylidene cyclohexanone analogs A-135 (81%), A-144 (58%), and A-154 (30%), synthesized efficiently via microwave-assisted green chemistry, exhibited superior multitarget binding affinities against key Mtb enzymes—A-135 (MtFabH, pantothenate synthetase), A-144 (MurE, DprE1, PTPs), and A-154 (Ddn, GlmU, Pks13)—with halogen substituents enhancing interactions through halogen bonds and lipophilicity; network pharmacology further revealed 294 overlapping TB-related targets and identified NF-κB1, STAT3, STAT1, and PTGS2 as key hubs mediating their multitarget therapeutic potential modulating TB clinical manifestation in human body.
Discussion
They exhibited superior multitarget binding affinities against key Mtb enzymes, A-135 (MtFabH, pantothenate synthetase), A-144 (MurE, DprE1, PTPs), and A-154 (Ddn, GlmU, Pks13)—with halogen substituents enhancing interactions through halogen bonds and lipophilicity; network pharmacology further revealed 294 overlapping TB-related targets and identified NF-κB1, STAT3, STAT1, and PTGS2 as key hubs mediating their multitarget therapeutic potential, modulating TB clinical manifestation in the human body.
Conclusion
A-135, A-144, and A-154 demonstrate promising multitarget anti-TB activity and potential as adjuvant therapies to complement existing treatments, especially for managing drug resistance and related comorbidities.
1. INTRODUCTION
Tuberculosis (TB), caused by Mycobacterium tuberculosis (Mtb), is transmitted via inhalation of aerosolized droplets and primarily infects alveolar macrophages, where it evades immune responses by inhibiting phagosome-lysosome fusion and neutralizing reactive oxygen species. These mechanisms promote intracellular survival, granuloma formation, and long-term latency, with reactivation often occurring under immunosuppressive conditions, leading to pulmonary cavitation and systemic spread [1-3]. Although initial infection is frequently asymptomatic, active TB typically presents with persistent cough, weight loss, fever, night sweats, and fatigue [3, 4]. Despite global health efforts, TB remains a major cause of mortality, with an estimated 1.25 million deaths in 2023, including 161,000 among HIV co-infected individuals. High-burden countries like India, Indonesia, China, and the Philippines account for over two-thirds of global cases, while diagnostic limitations and rising multidrug-resistant TB (MDR-TB), for which only a small fraction receive effective treatment, continue to undermine disease control efforts [4].
The complex pathology and resistance potential of TB necessitate multifaceted drug regimens targeting heterogeneous bacterial populations. For drug-susceptible tuberculosis (TB), the World Health Organization (WHO) advises a six-month, combined treatment regimen incorporating isoniazid, rifampin, pyrazinamide, and ethambutol. Conversely, multidrug-resistant TB (MDR-TB) necessitates the utilization of novel, all-oral treatment protocols. These treatment regimens incorporate agents, such as bedaquiline, pretomanid, linezolid, and moxifloxacin, with the objectives of enhancing patient compliance, shortening the duration of therapy, and increasing cure rates. However, the prolonged use of several antibiotics often results in adverse effects. These encompass hepatotoxicity, peripheral neuropathy, and gastrointestinal disturbances, which can compromise patient adherence, contribute to treatment discontinuation, and promote the emergence of resistance [5].
Persistent challenges in tuberculosis (TB) treatment, such as extended treatment regimens, negative drug effects, and the increasing incidence of multidrug resistance, have prompted research into natural bioactive compounds as adjunct therapeutic agents. Curcumin, a polyphenolic substance sourced from Curcuma longa, exhibits a broad spectrum of pharmacological characteristics, including antioxidant, anti-inflammatory, anti-microbial, and hepatoprotective properties [6-9]. Moreover, its immuno-modulatory function, particularly its ability to enhance T-cell responses, highlights its potential therapeutic value [10]. Conversely, the clinical use of curcumin has been historically limited by its poor bioavailability, which has led to the development of nanoparticle-based formulations.
These formulations significantly enhance curcumin's solubility, stability, and systemic absorption [11, 12]. Preclinical studies indicate that nano-curcumin can improve the efficacy of standard anti-TB drugs, mitigate hepatotoxicity, and strengthen host immune responses. Consequently, this suggests its ability to improve treatment adherence, shorten the duration of therapy, and decrease both relapse and resistance, thus underscoring its potential as a multifunctional adjuvant in TB treatment [6].
Recent progress in the development of tuberculosis (TB) therapeutics has increasingly focused on curcumin analogs, which are structurally altered derivatives designed to improve chemical stability, biological efficacy, and target specificity. Monocarbonyl curcumin analogs, which substitute the native β-diketone moiety with a monoketone group, have demonstrated enhanced chemical resilience and bioactivity, displaying notable anti-inflammatory, anti-diabetic, and antitubercular effects in both in vitro and in vivo investigations [13]. Furthermore, structural modifications, such as ring-closing alterations and heterocyclic substitutions, have expanded their therapeutic applications. Simultaneously, the deliberate halogenation of established anti-TB agents, including fluorinated derivatives of p-aminosalicylic acid and thioacetazone, has proven effective in augmenting antimycobacterial activity.
These halogenated compounds are being increasingly used in combination therapies, especially for treating multidrug-resistant strains of tuberculosis. Specifically, the addition of fluorine improves how the drug is absorbed and distributed in the body, how easily it enters cells, and how well it binds to its target. This leads to a stronger antimicrobial effect overall [14, 15]. Moreover, when these halogenated agents are used with new treatments like bedaquiline, delamanid, or pretomanid, they work together, which can shorten treatment times, delay the development of drug resistance, and improve patient outcomes in cases of resistant TB.
Alongside progress in pharmacology, drug development is progressively oriented towards synthetic methodologies that prioritize both efficiency and environmental sustainability. While traditional synthetic routes remain widespread, they often encounter difficulties, including protracted reaction times, stringent conditions, and inadequate product yields [16]. To mitigate these limitations, microwave-assisted synthesis has emerged as a promising green chemistry approach, utilizing electromagnetic irradiation to facilitate rapid and uniform heating through dipolar polarization and ionic conduction.
Microwave-assisted synthesis significantly accelerates reaction kinetics, resulting in reaction rate enhancements that are considerably superior to those obtained via conventional approaches. This methodology improves reaction efficiency, enhances product purity, and reduces by-products and waste, often operating under solvent-free or environmentally friendly conditions. By selectively heating reactants rather than the entire system, microwave synthesis diminishes energy consumption and operational risks, thus positioning itself as a crucial tool for sustainable, reproducible, and safe synthetic procedures [17].
While statistical power calculations were not performed, the selection of the number of compounds was informed by prior exploratory docking studies of a comparable nature within the framework of early drug discovery. MurE, Ddn, GlmU, Pks13, DprE1, MtFabH, PanC, and PtpA were chosen as key targets for peptidoglycan biosynthesis and metabolism in Mycobacterium tuberculosis (Mtb), given their essential functions in tuberculosis (TB) pathogenesis. Therefore, lead compounds with strong binding affinities were subjected to network pharmacology analyses to predict their interactions with host-pathogen pathways, thereby confirming their potential as immunomodulatory agents for improving TB treatment. To ensure methodological consistency with the study's goals, an exploratory, non-clinical in silico design was used, incorporating a quantitative methodology based on synthetic compound assessment and bioinformatics analysis. The literature supporting target selection and compound validation was gathered through structured searches of the PubMed, Scopus, and Web of Science databases, using predefined keywords and inclusion criteria that focused on curcumin derivatives and TB-related targets published between 2015 and 2025. Protein and gene targets were obtained from UniProt, GeneCards, SwissTargetPrediction, SuperPred Target Prediction, and NCBI. Interaction networks and pathway enrichment analyses were performed using the STRING database and Cytoscape software; concurrently, gene ontology (GO) analysis was conducted via the WebGestalt server. This integrated approach facilitated the systematic identification and assessment of multitarget interactions, alongside the host-modulatory potential of the analogues, thereby informing the rational design of potential anti-TB agents.
2. MATERIALS AND METHODS
2.1. Chemicals, Reagents, and Instrumentation
The series of curcumin analog compounds A-125, A-128, A-135, A-137, A-143, A-144, A-150, A-152, and A-154 were synthesized at the Curcumin Research Center (CRC), Prof. Ritmaleni’s Laboratory, Faculty of Pharmacy, Universitas Gadjah Mada, Indonesia. Microwave-assisted synthesis was executed utilizing a Monowave 400 microwave reactor, supplied by Anton Paar, an Austrian company. Aldehydes were sourced from Sigma Aldrich, a United States-based entity, whereas cyclohexanone, benzaldehydes, and thin-layer chromatography (TLC) precoated silica gel plates (GF256 type 60) were procured from Merck, a German corporation. Furthermore, a variety of organic solvents, encompassing glacial acetic acid, ethanol, acetone, and water, were obtained from Sigma Aldrich. The synthesized compounds underwent characterization via Thin-Layer Chromatography (TLC), Infrared (IR) Spectroscopy, Nuclear Magnetic Resonance (NMR) Spectroscopy, and High-Resolution Mass Spectrometry (HRMS). The detailed synthetic methodologies for each compound are detailed in the synthesis procedure section and the Supplementary Materials.
2.2. Synthesis of Curcumin Analogues via Microwave- Assisted Method
Microwave irradiation utilizes electromagnetic waves to uniformly and swiftly heat reactants, thus promoting condensation reactions, exemplified by the synthesis of benzylidene-cyclohexanone; this methodology further reduces energy expenditure, reaction time, and the use of hazardous substances, thereby adhering to green chemistry principles. The synthesis of benzylidene-cyclohexanone compounds, especially benzylidene-cyclohexanone derivatives, typically involves a Claisen-Schmidt condensation. This procedure necessitates the reaction of substituted benzaldehydes (2 molar equivalents) with cyclohexanone (1 molar equivalent) in the presence of an acid catalyst (HCl), sometimes alongside glacial acetic acid, within a suitable organic solvent. The reactions were performed using a Monowave 400 microwave synthesizer (Anton Paar), where the reaction mixtures were stirred at 600 rpm and irradiated at 80°C for roughly 9 minutes.
Following this, the crude products underwent separation via solvent washing, commonly utilizing ethanol-water or acetone-water systems, and were subsequently purified through recrystallization, resulting in the ultimate yellow crystalline solids. For the synthesis of particular compounds, including 2,6-bis(2’-chlorobenzylidene) cyclohexanone (A-125), 2,6-bis(2’-bromobenzylidene) cyclohexanone (A-128), or 2,6-bis(4’-chlorobenzylidene) cyclohexanone (A-144), substituted benzaldehydes, such as 2-chlorobenzaldehyde, 2-bromobenzaldehyde, 3-bromobenzaldehyde, or 4-chlorobenzaldehyde, were employed for each derivative.
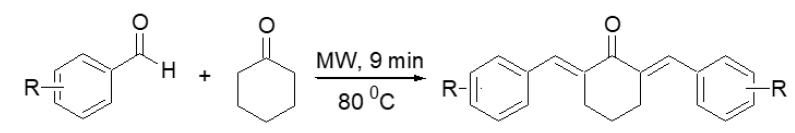
Synthesis of benzylidene-cyclohexanone analogues by microwave irradiation.
After microwave irradiation, the products were extracted, washed, and recrystallized using solvent mixtures chosen based on the solubility and purity characteristics of each compound. Subsequently, the final products were characterized via Thin-Layer Chromatography (TLC), melting point determination, Infrared (IR) Spectroscopy, Nuclear Magnetic Resonance (NMR) Spectroscopy, and High-Resolution Mass Spectrometry (HRMS), which confirmed their structural integrity and purity, as illustrated in Scheme 1.
2.3. Molecular Docking Studies
Molecular docking analysis was performed using Molecular Operating Environment (MOE) version 2015 (licensed by the Department of Pharmaceutical Chemistry, Faculty of Pharmacy, Universitas Gadjah Mada) to investigate interactions with several essential proteins of Mtb.
The literature supporting docking target selection was obtained through unsystematic exploration of relevant articles across PubMed, Scopus, and Web of Science. To identify relevant studies, keywords such as “tuberculosis,” “curcumin analogues,” “molecular docking,” “multitarget therapy,” and “network pharmacology” were used. Articles published between 2015 and 2025, written in English, and discussing TB-related targets or curcumin-based compounds were included. Peer-reviewed articles with experimental or computational findings were prioritized, while reviews without supporting data, case reports, and sources not indexed in databases were usually excluded. These targets, retrieved from the Protein Data Bank (PDB) via the RCSB website (https://www.rcsb.org), include MurE (PDB ID: 2WTZ), a ligase involved in peptidoglycan synthesis; Ddn (PDB ID: 3R5L), a nitroreductase activating prodrugs; GlmU (4G87), a key enzyme in cell wall precursor biosynthesis; Pks13 (PDB ID: 5V3Y), a polyketide synthase critical for mycolic acid production; DprE1 (PDB ID: 6G83), essential for arabinogalactan synthesis; MtFabH (PDB ID: 2QX1), fatty acid synthase III; Pantothenate Synthetase, involved in coenzyme A biosynthesis; and PTP A (PDB ID: 1U2Q), a virulence factor modulating host immune responses. Protein preparation involved removing water, solvents, and cholesterol. Then, hydrogen atoms were added using the “Protonate 3D” feature [18-20]. Energy minimization was performed using the MOE “Energy Minimization” tool with the default settings. The docking protocol was validated by finding the binding site with the “Surface and Maps” tool and redocking. Results were considered acceptable if the root-mean-square deviation (RMSD) was less than 2 Å. The 2D structures of halogenated benzylidene curcumin analogues, along with the reference drugs isoniazid (isonicotinic acid hydrazide/H), pyrazinamide (Z), and ethambutol (E), were converted to SMILES format. These were then transformed into 3D conformations using the “Builder” function.
The MOE database was created by assembling these compounds, assigning partial charges, and minimizing their energy. Docking simulations were performed, using the binding site defined by “Ligand Atoms,” the “Triangle Matcher” method for placement, and “Induced Fit” for refinement. The scoring functions used were “London dG” as the main scoring parameter and “GBVI/WSA dG” as the secondary. The conformation showing the lowest binding affinity was selected for further analysis and visualized in both 2D and 3D formats.
2.4. Network Pharmacology Analysis
A comprehensive bioinformatics strategy was applied in this study to predict potential protein targets of the combined halogenated compounds A-135, A-144, and A-154, utilizing multiple publicly available target prediction platforms. The network pharmacology approach used in this work was adapted from previous studies, notably the publication “Identification of potential targets of the curcumin analog CCA-1.1 for glioblastoma treatment: integrated computational analysis and in vitro study” [21]. That study successfully integrated network pharmacology with experimental validation, demonstrating the utility of this approach in early-stage drug discovery. Given the favorable correlation between in silico predictions and laboratory findings reported therein, the adopted methodology is considered reliable for preliminary target identification and hypothesis generation in this study. Target prediction and target fishing were employed to identify genes linked to tuberculosis (TB) infection and its significant clinical outcomes, encompassing malnutrition, anemia, inflammatory responses associated with Mycobacterium tuberculosis (Mtb) genes, and multidrug-resistant tuberculosis (MDR-TB). These investigations leveraged databases, such as SwissTargetPrediction (http://www.swisstargetprediction.ch), SuperPred Target Prediction (https://prediction.charite.de/subpages/target_ prediction.php), GeneCards (https://www.genecards.org/#), and NCBI (https://www.ncbi.nlm.nih.gov/) to ascertain potential protein targets implicated in TB modulation, designated as THTGTs (Tuberculosis and Halogenated-Benzylidene Target Genes) [22-27]. Overlapping gene targets were determined via InteractiVenn (https://www. interactivenn.net). Subsequently, protein-protein interaction (PPI) networks were constructed using the STRING database version 11.5 to investigate functional relationships among the identified proteins. Subsequently, the network data underwent analysis via Cytoscape software, specifically version 3.9.1, with the CytoHubba plugin utilized to ascertain the ten most critical hub genes, as assessed by the “Degree Score” algorithm [28, 29].
Moreover, to clarify the biological relevance of these hub genes, functional enrichment analysis was performed. Gene Ontology (GO) analysis was conducted using the WEB-based Gene Set Analysis Toolkit (WebGestalt) (https://webgestalt.org/), employing the Over-Representation Analysis (ORA) methodology. The analysis was set to Homo sapiens as the reference organism, and results were filtered by applying a false discovery rate (FDR) threshold (< 0.05) to maintain statistical validity [30].
3. RESULTS AND DISCUSSION
3.1. Chemical Synthesis
Prior investigations have demonstrated that halogenated analogs of anti-TB medications can provide more efficacious treatment for Mycobacterium tuberculosis infections, thereby augmenting therapeutic efficacy relative to their non-halogenated counterparts. Moreover, these derivatives frequently exhibit reduced toxicity, consequently diminishing the probability of adverse patient reactions. This attribute renders halogenated derivatives appealing candidates for the development of safer therapeutic alternatives. Furthermore, halogenated compounds have been observed to possess enhanced pharmacokinetic characteristics, such as improved bioavailability and extended half-lives, which can further contribute to improved clinical results.
In the synthetic procedures undertaken, compound A-135 yielded the highest output, as illustrated in Fig. (1) and Table 1, achieving a yield of 81%, whereas compound A-152 demonstrated the lowest yield, at a mere 23%. An examination of the substitution patterns uncovers noteworthy trends concerning the influence of halogen type and position on the observed yields. Chlorine, when situated at the ortho position—as demonstrated in entries 1 and 6—frequently yielded superior results compared to its placement at the para position. Furthermore, the incorporation of chlorine in the ortho position (entries 1 and 2) yielded a greater output than bromine in a similar configuration, thereby implying a positional advantage for chlorine over bromine within this specific framework. Conversely, bromine at the meta position, as evidenced in entries 2 and 3, produced higher yields than when positioned at the ortho site, suggesting that bromine is favored by a meta placement. This trend persisted. When bromine was positioned meta, as in entries 3 and 4, the results were consistently superior. Specifically, bromine at the para position exhibited better performance than chlorine at the same site (entries 4 and 6), whereas chlorine at the para position outperformed fluorine when both were similarly positioned (entries 6 and 8). Furthermore, when both chlorine and fluorine were located at the ortho position, as shown in entries 5 and 7, the compound with two chlorine atoms yielded results comparable to those of compounds containing both halogens. Comparisons between ortho and para substitutions also indicated that chlorine in the ortho position, as in entries 1 and 5, generally produced higher yields than in the para position. However, the reverse was observed in entries 1 and 7, where para-substituted chlorine resulted in better yields. Finally, entries 7 and 9 demonstrated that chlorine in the ortho and para positions yielded comparable results.
Microwave irradiation facilitates highly effective energy transfer, thereby accelerating molecular activation and enhancing reaction kinetics. Consequently, reaction durations are reduced to minutes, as opposed to hours, which improves laboratory efficiency and decreases solvent waste. Furthermore, the uniform heating profile diminishes the generation of byproducts commonly arising from localized overheating in traditional methodologies. Regarding the synthesis of benzylidene-cyclohexanone, this approach not only achieves favorable to excellent product recovery (23–81% yields) but also allows for the scalable production of a diverse array of derivatives featuring various aromatic substitutions, thus enabling further investigation into their chemical and biological characteristics.
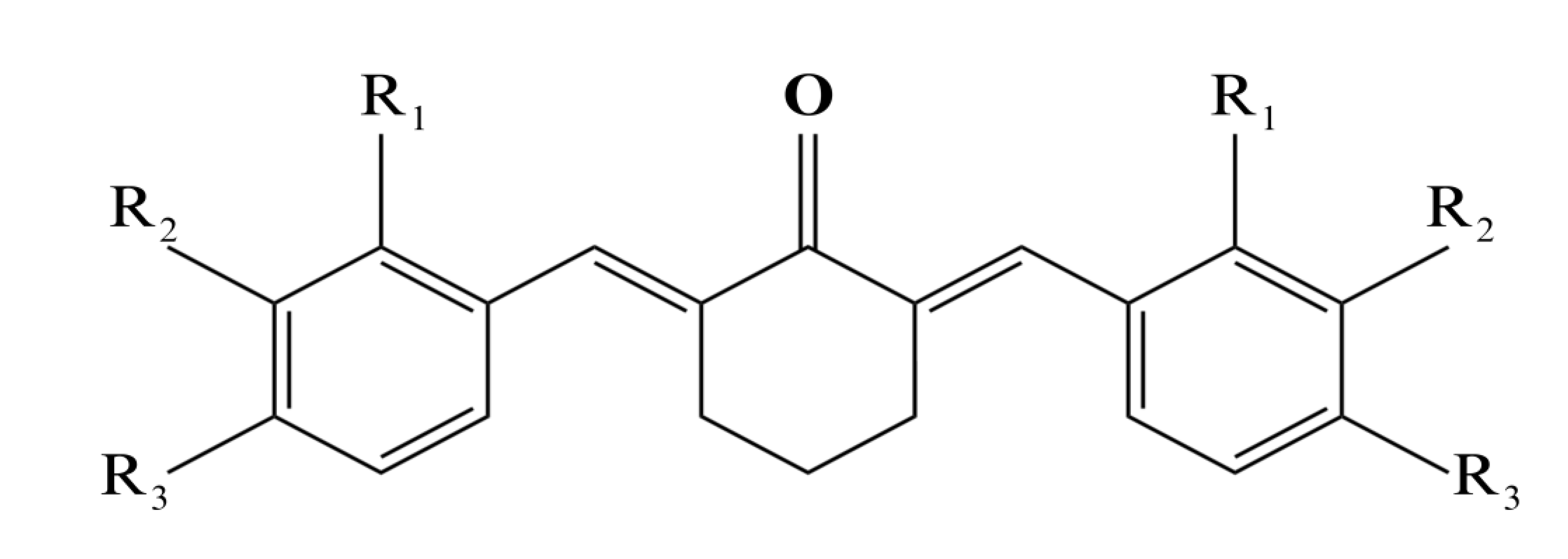
Main backbone chain of halogenated benzylidene cyclohexanone.
| Entry | R-Benzaldehyde | Code | R1 | R2 | R3 | R4 | R5 | Yield (%) |
|---|---|---|---|---|---|---|---|---|
| 1 | 2-Chlorobenzaldehyde | A-125 | Cl | H | H | H | H | 67 |
| 2 | 2-Bromobenzaldehyde | A-128 | Br | H | H | H | H | 56 |
| 3 | 3-Bromobenzaldehyde | A-135 | H | Br | H | H | H | 81 |
| 4 | Bromobenzaldehyde | A-137 | H | H | B | H | H | 74 |
| 5 | 2-Chloro-6-fluorobenzaldehyde | A-143 | Cl | H | H | H | F | 29 |
| 6 | 4-Chlorobenzaldehyde | A-144 | H | H | Cl | H | H | 58 |
| 7 | 2,6-Dichlorobenzaldehyde | A-150 | Cl | H | H | H | Cl | 29 |
| 8 | 4-Fluorobenzaldehyde | A-152 | H | H | F | H | H | 23 |
| 9 | 2,6-Dichlorobenzaldehyde | A-154 | Cl | H | Cl | H | H | 30 |
3.2. Molecular Docking
Molecular docking multitarget assays on Mtb receptors reveal that curcumin analogs, specifically benzylidene cyclohexanone derivatives with halogen substituents, as shown in Fig. (2) and Table 2, demonstrate promising potential as novel anti-TB agents. Compound A-144 exhibited the highest binding affinity, as evidenced by the most negative docking scores against MurE (-7.21 kJ/mol), DprE1 (-7.6 kJ/mol), and phosphate acetyltransferase (PTP, -5.68 kJ/mol). Compound A-154 was determined to be the most potent inhibitor of Ddn (-5.3 kJ/mol), GlmU (-5.31 kJ/mol), and Pks13 (-6.65 kJ/mol), while compound A-128 displayed the strongest interactions with MtFabH (-6.96 kJ/mol) and pantothenate synthetase (-5.68 kJ/mol). These binding affinities surpass those of established anti-TB drugs, such as isoniazid, pyrazinamide, and ethambutol. Therefore, a combination of A-135, A-144, and A-154 could potentially represent a strong multitarget approach to TB treatment, resulting in improved antimicrobial effectiveness.
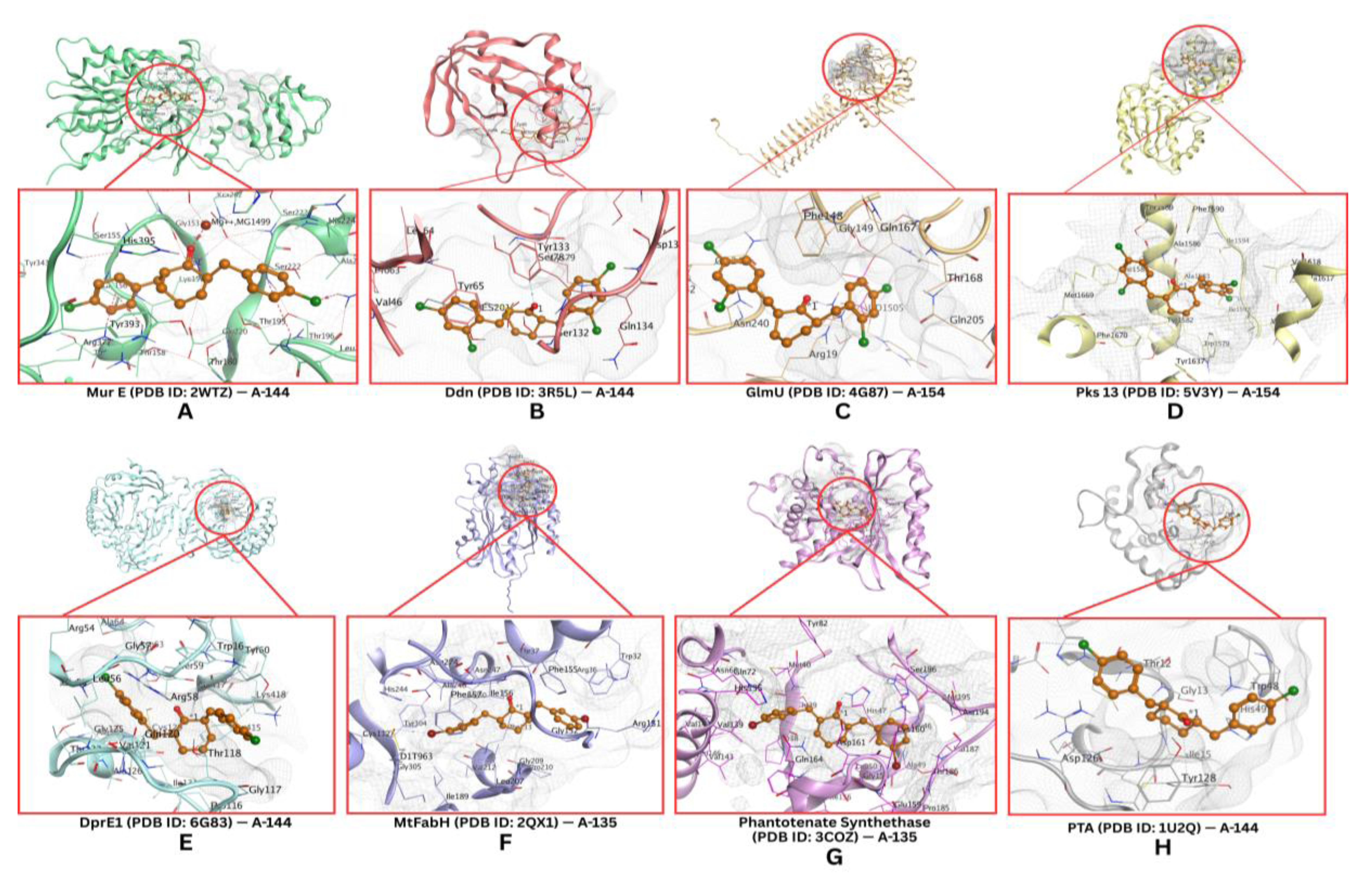
The molecular docking results of halogenated benzylidene-cyclohexanone compounds against Mycobacterium tuberculosis (Mtb) receptors showed the highest affinities for compound A-135 with MtFabH and pantothenate synthetase; compound A-144 with MurE, DprE1, and PTP; and compound A-155 with Ddn, GlmU, and Pks1.
| Mycobacterium Tuberculosis Targets | A-125 | A-128 | A-135 | A-137 | A-143 | A-144 | A-150 | A-152 | A-154 | H | Z | E | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mur E (2WTZ) | Dock. Score | -6.86 | -6.69 | -6.12 | -6.84 | -6.78 | -7.21 | -6.18 | -6.57 | -6.43 | -4.79 | -4.64 | -5.77 |
| Residues | Ser 155, Gly 156, Lys 157, Thr 159, Glu 220, Arg 230, Arg 377, Asp 392, Tyr 393, Lys 396 | ||||||||||||
| Ddn (3R5L) | Dock. Score | -4.53 | -4.80 | -5.04 | -5.13 | -4.50 | -4.86 | -4.63 | -4.68 | -5.30 | -3.91 | -4.92 | -4.00 |
| Residues | Tyr 65, Ser 78, Lys 79, Gly 89, Ser 132, Asp 135 | ||||||||||||
| GlmU (4G87) | Dock. Score | -5.13 | -4.61 | -4.96 | -5.00 | -4.71 | -4.79 | -4.31 | -4.38 | -5.31 | -6.33 | -7.04 | -5.29 |
| Residues | Arg 19, Thr 168, Gln 205, Glu 207 | ||||||||||||
| pks 13 (5V3Y) | Dock. Score | -5.6 | -5.55 | -5.54 | -5.42 | -5.91 | -5.7 | -5.92 | -5.73 | -6.65 | -4.23 | -4.48 | -4.17 |
| Residues | Ile 1507, Tyr 1582, Val 1614, Gln 1633, Thr 1645 | ||||||||||||
| DprE1 (6g83) | Dock. Score | -7.55 | -7.37 | -7.54 | -7.3 | -6.8 | -7.6 | -6.55 | -7.52 | -7.51 | -5.05 | -6.59 | -4.5 |
| Residues | Leu 56, Tyr 60, Thr 122, Gly 182, Ile 184 | ||||||||||||
| MtFabH (2qx1) | Dock. Score | -6.17 | -6.64 | -6.96 | -6.75 | -6.1 | -6.6 | -6.84 | -6.66 | -6.76 | -4.22 | -5.12 | -4.59 |
| Residues | Asp 27, Trp 32, Arg 151, Gly 152, Gly 209, Asn 274, Tyr 304 | ||||||||||||
| Phantotenate Synthethase | Dock. Score | -7.38 | -7.01 | -8.17 | -7.23 | -6.6 | -7.66 | -4.91 | -7.68 | -6.75 | -4.81 | -5.63 | -4.08 |
| Residues | Pro 38, His 44, Phe 156, Gly 158, Gln 164, Met 195 | ||||||||||||
| PTP (1u2q) | Dock. Score | -5.47 | -5.41 | -5.46 | -5.48 | -5.39 | -5.68 | -5.53 | -5.36 | -5.56 | -4.39 | -5.61 | -4.51 |
| Residues | Thr 12, Ile 15, Arg 17, Trp 48, Asn 92 | ||||||||||||
Compound A-135's mode of action entails the suppression of MtFabH and pantothenate synthetase, both of which are essential enzymes within Mycobacterium tuberculosis's (Mtb) lipid metabolism and energy production pathways. MtFabH, also known as β-ketoacyl-ACP synthase III, plays a pivotal role in the commencement of fatty acid elongation, a process critical for the biosynthesis of mycolic acids. These mycolic acids are indispensable lipid components of the Mtb cell wall, providing structural stability and resistance to both host immune defenses and antibiotic interventions. As a result, the inhibition of MtFabH activity obstructs the synthesis of these protective lipids, thus increasing the bacterium's vulnerability to external threats. Furthermore, pantothenate synthetase facilitates the reaction between pantoate and β-alanine, yielding pantothenate, also known as vitamin B5. This vitamin serves as a precursor to coenzyme A (CoA), a critical component in fatty acid synthesis, energy metabolism, and numerous other biosynthetic pathways. Consequently, by inhibiting both MtFabH and pantothenate synthetase, A-128 concurrently compromises the bacterial envelope and impedes the cell's acquisition of the metabolic resources necessary for growth and survival. This dual action effectively prevents Mtb from maintaining its viability [31, 32].
Compound A-144 exerts its effects by inhibiting MurE, DprE1, and PTP, thereby interfering with the construction and upkeep of the Mycobacterium tuberculosis (Mtb) cell wall and its fundamental metabolic functions. MurE, an indispensable ligase within the peptidoglycan biosynthesis pathway, facilitates the incorporation of meso-diaminopimelic acid into the elongating stem peptide, thereby ensuring the cross-linking that confers mechanical stability to bacterial cell walls. Conversely, DprE1 functions as a crucial epimerase in the synthesis of arabinogalactan, a polysaccharide that connects the peptidoglycan layer to the mycolic acid-rich outer membrane, which is a characteristic of the Mtb cell wall. Protein tyrosine phosphatases (PTPs) in Mycobacterium tuberculosis (Mtb) function as signaling enzymes, modulating phosphorylation-dependent signaling pathways that govern cell division, stress responses, and virulence. The inhibition of PTPs compromises the bacterium's capacity to orchestrate critical cellular processes, thereby diminishing its capacity to adapt and survive in adverse environments. A-144, by concurrently targeting these three elements, disrupts the bacterium's capacity to produce a stable cell wall and induces a significant metabolic impediment, which in turn leads to the failure of vital biosynthetic and energetic pathways, ultimately resulting in bacterial death [33-35].
Compound A-154, the most promising inhibitor of Ddn, GlmU, and Pks13, affects several key processes that are essential for Mycobacterium tuberculosis (Mtb) to survive stress and build its outer membrane. Ddn (deazaflavin-dependent nitroreductase) is crucial for activating nitroimidazole prodrugs and managing nitrosative stress, a defense mechanism Mtb uses to survive the hostile environment inside host macrophages. As a result, inhibiting Ddn weakens the bacterium's ability to resist oxidative and nitrosative stress, making it more vulnerable to immune system attacks. GlmU, a bifunctional enzyme, facilitates the concluding stages of UDP-N-acetylglucosamine synthesis, which is essential for peptidoglycan construction. Conversely, Pks13 is responsible for the ultimate condensation of fatty acyl chains, thereby generating mycolic acids, which are crucial constituents of the mycobacterial outer membrane. A-154's simultaneous targeting of Ddn, GlmU, and Pks13 compromises the bacterium's stress response, inhibits the provision of cell wall precursors, and impedes mycolic acid biosynthesis, thereby directly affecting Mtb's structural integrity and its survival mechanisms [36-38].
The presence of chlorine (Cl) and bromine (Br) substituents in these compounds is crucial for their biological activity. These halogen atoms augment the molecules' lipophilicity, thereby improving membrane permeability and fostering more robust interactions within the hydrophobic regions of their target enzymes. The greater atomic radius and polarizability of bromine in A-135 enable the establishment of more significant van der Waals and hydrophobic interactions. Conversely, the chlorine atoms in A-144 and A-154 exert electron-withdrawing effects, which stabilize halogen bonding and intensify electrostatic and dipole interactions at the binding sites. Furthermore, the inclusion of two chlorine substituents in A-154 may facilitate improved spatial fitting within the enzyme active sites, consequently increasing its affinity and specificity. Collectively, the incorporation of these halogens fine-tunes the compounds’ binding profiles and improves their overall potency as multitarget anti-TB agents [39, 40]. The enhanced binding affinities exhibited by these curcumin derivatives, in comparison to existing tuberculosis (TB) therapies, such as HZE, highlight their promise as a new category of multitarget anti-TB agents. The combination of A-128, A-144, and A-154, which collectively engage with multiple critical enzymatic systems, implies the potential for heightened antimicrobial effectiveness and a reduced likelihood of resistance development. Moreover, the benzylidene cyclohexanone compounds that displayed the most robust affinity for Mycobacterium tuberculosis (Mtb) targets were those produced with high yields.
Employing a microwave-assisted, environmentally conscious synthetic strategy, we synthesized A-135 (81%) and A-144 (58%). Despite A-154's lower yield of 30% relative to the other compounds, this outcome underscores the microwave-assisted synthesis method's utility in drug discovery, especially for potential anti-TB agents, while simultaneously adhering to green chemistry principles. Although the remaining analogs were not extensively characterized, they exhibited encouraging biological activity, with binding affinities typically surpassing those of existing anti-Mtb drugs (HZE), thereby indicating the necessity for further investigation and development.
3.3. Network Pharmacology
This investigation utilized a thorough bioinformatics methodology, specifically network pharmacology, to clarify the intricate relationships between tuberculosis (TB)-associated genes and the therapeutic efficacy of halogenated benzylidene curcumin analogues A-135, A-144, and A-154. This approach facilitated the identification of novel therapeutic targets, pertinent biological pathways, and potential hub genes, with a particular focus on those associated with significant clinical manifestations of TB, thus providing crucial information for prospective therapeutic advancements. Malnutrition, anemia, and inflammatory responses were highlighted among these manifestations, given their essential roles in disease progression, treatment outcomes, and clinical prognosis. Malnutrition, for instance, undermines host immunity and elevates the risk of active TB by compromising immune defense mechanisms [41]. Anemia frequently manifests in individuals afflicted with tuberculosis (TB), and its presence correlates with the severity of the disease; this association may be attributable to chronic inflammation and the dysregulation of iron metabolism, potentially mediated by the hepcidin pathway [42, 43]. Moreover, the inflammatory responses instigated by Mycobacterium tuberculosis (Mtb) can precipitate significant tissue damage and sustained immune activation, thereby intensifying the pathological consequences of the disease [44]. The emphasis on multidrug-resistant TB (MDR-TB) is warranted, given its escalating prevalence and the associated clinical difficulties, which encompass restricted therapeutic alternatives and elevated mortality rates [45].
Considering the clinical relevance of the identified phenomena, they were incorporated into the network pharmacology analysis, functioning as crucial reference points for target mapping and pathway enrichment. Target gene prediction was performed using SwissTarget Prediction, SuperPred, GeneCards, and NCBI databases to identify genes associated with tuberculosis and its associated comorbidities. This approach enabled the identification of shared genes affected by the halogenated compounds, thus supporting their potential utility in the modulation of various aspects of tuberculosis pathology.
Based on predictions from SwissTargetPrediction, the combination of A-135, A-144, and A-154 was associated with 300 potential molecular targets (Supplementary Table 1). In parallel, the SuperPred Target Prediction platform identified 370 receptor-level targets for the same compound set (Supplementary Tables 2 and 3). GeneCards and NCBI provided a detailed compilation of 19,587 genes linked to significant clinical features of TBB, encompassing malnutrition, anemia, inflammatory reactions induced by Mtb, and multidrug-resistant TB (MDR-TB) (Supplementary Table 4). An integrative analysis employing a Venn diagram identified 294 shared genes, designated as Tuberculosis and Halogenated-Benzylidene Target Genes (THTGTs), which were then chosen for subsequent functional and network analyses (Fig. 3A-D).
The PPI network illustrating the interconnections among the proteins is shown in Fig. (3C) (Supplementary Tables 5 and 6). The hub genes associated with this compound were identified, with the top 10 genes based on degree score: EGFR, HSP90AA1, STAT3, PTGS2, HSP90AB1, NF-κB1, EP300, APP, STA1, and NR3C1 (Fig. 3D). Bioinformatic analysis shows that the combination of these three compounds holds significant potential in overcoming TB and its clinical manifestation through 4 principal proteins identified on high degree score: NF-κB1, STAT3, STAT1, and PTGS2. These proteins play a significant role in regulation and their relevance to TB-associated clinical manifestations (Fig. 4).
To clarify the potential mechanisms and therapeutic importance of the curcumin analogues identified in this study (A-135, A-144, and A-154), we performed Gene Ontology (GO) enrichment analysis. This analysis covered Biological Process (BP), Molecular Function (MF), and Cellular Component (CC) categories. GO analysis is a key bioinformatic method for understanding the functions of target genes and their roles in disease-related pathways, particularly in host-pathogen interactions, such as those seen in tuberculosis (TB) (Fig. 3B) [46, 47].
Gene Ontology (GO) enrichment analysis was performed to elucidate the functional profiles of predicted target genes influenced by halogenated curcumin analogs A-135, A-144, and A-154. WebGestalt's GO visualization commonly employs bar plots; the x-axis displays enriched GO terms, ordered by statistical significance, while the y-axis indicates the number of genes linked to each term. A greater gene count signifies a more substantial enrichment and suggests a higher biological significance of the term within the analyzed dataset [30].
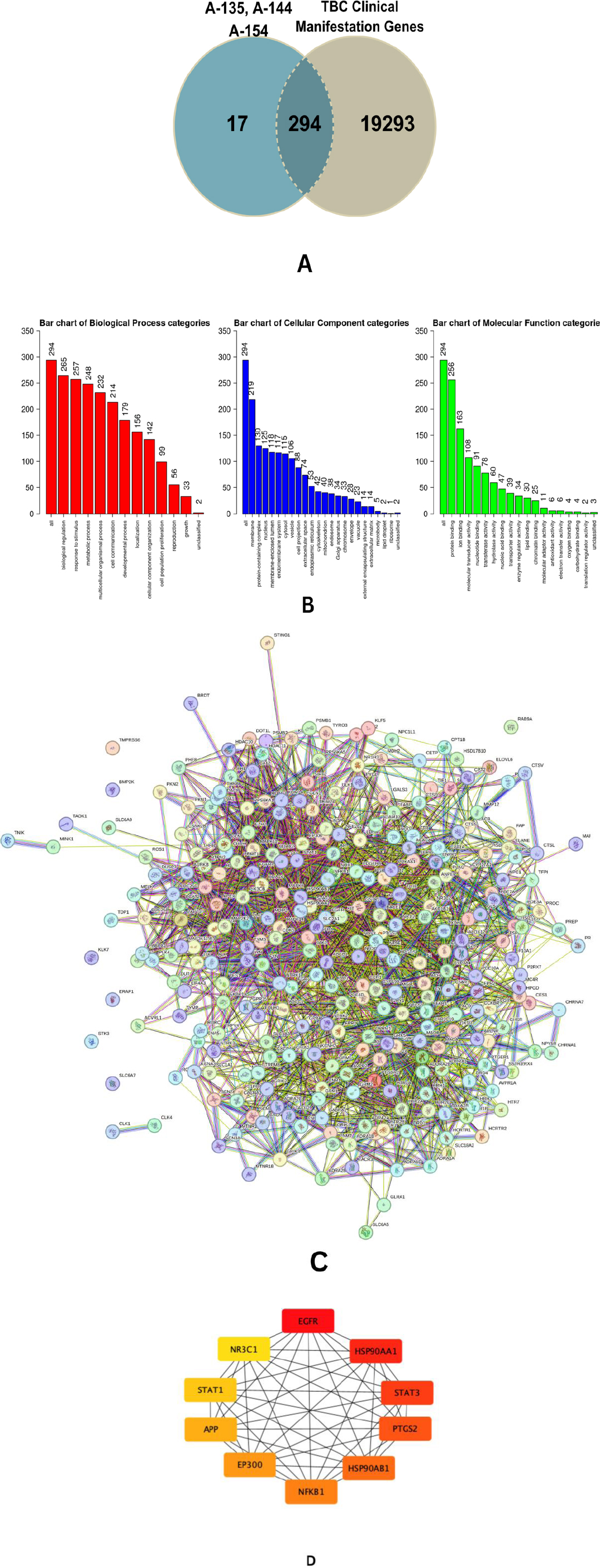
(A) Venn diagram showing 294 predicted target genes identified as THTGTs. (B) GO enrichment analysis of PTPMs categorized into Biological Process (BP), Cellular Component (CC), and Molecular Function (MF). (C) Protein–protein interaction (PPI) network of THTGTs constructed using STRING, consisting of 293 nodes and 2,324 edges. The average local clustering coefficient, which is 0.405, indicates a high level of interconnectedness among the targets. (D) The ten most important hub genes, ranked by degree score, were identified using the CytoHubba plugin in Cytoscape. The color of each node represents its degree score, with red indicating the highest scores, followed by orange and yellow.
Under the Biological Process (BP) category, the most significantly enriched terms included biological regulation (265 genes), response to stimulus (257 genes), and metabolic processes (248 genes). The high number of genes involved in biological regulation suggests that these compounds may influence core regulatory networks, including transcriptional control, cytokine signaling, and homeostatic adaptation—processes frequently subverted during Mtb infection. Similarly, enrichment in response to stimulus indicates potential roles in modulating host defense mechanisms and cellular stress pathways, while the metabolic process enrichment suggests involvement in altering bioenergetic and metabolic reprogramming, a key survival strategy of intracellular pathogens. Within the Molecular Function (MF) domain, top-enriched terms included protein binding (256 genes), ion binding (163 genes), and molecular transducer activity (108 genes). The pattern implies that the target compounds may modulate protein–protein interactions and receptor-mediated signaling, consistent with their predicted engagement with immune-regulatory proteins, such as STAT1 and PTGS2 (COX-2). These molecular functions are essential for transmitting extracellular signals to intracellular responses during infection and inflammation. Cellular Component (CC) analysis showed that gene products were mainly found in the membrane (219 genes), protein-containing complexes (130 genes), and the nucleus (125 genes). This pattern of localization supports the idea of multiple compartments being involved in how these genes work. This includes activating membrane-bound receptors, like toll-like receptors, forming intracellular complexes, such as inflammasomes, and regulating transcription in the nucleus. The presence of PTGS2 and STAT1 in these compartments also supports their potential as important molecular targets. Overall, these Gene Ontology (GO) enrichment results highlight the curcumin analogs' potential to both fight microbes and modulate the immune system in interactions between hosts and pathogens [48, 49].
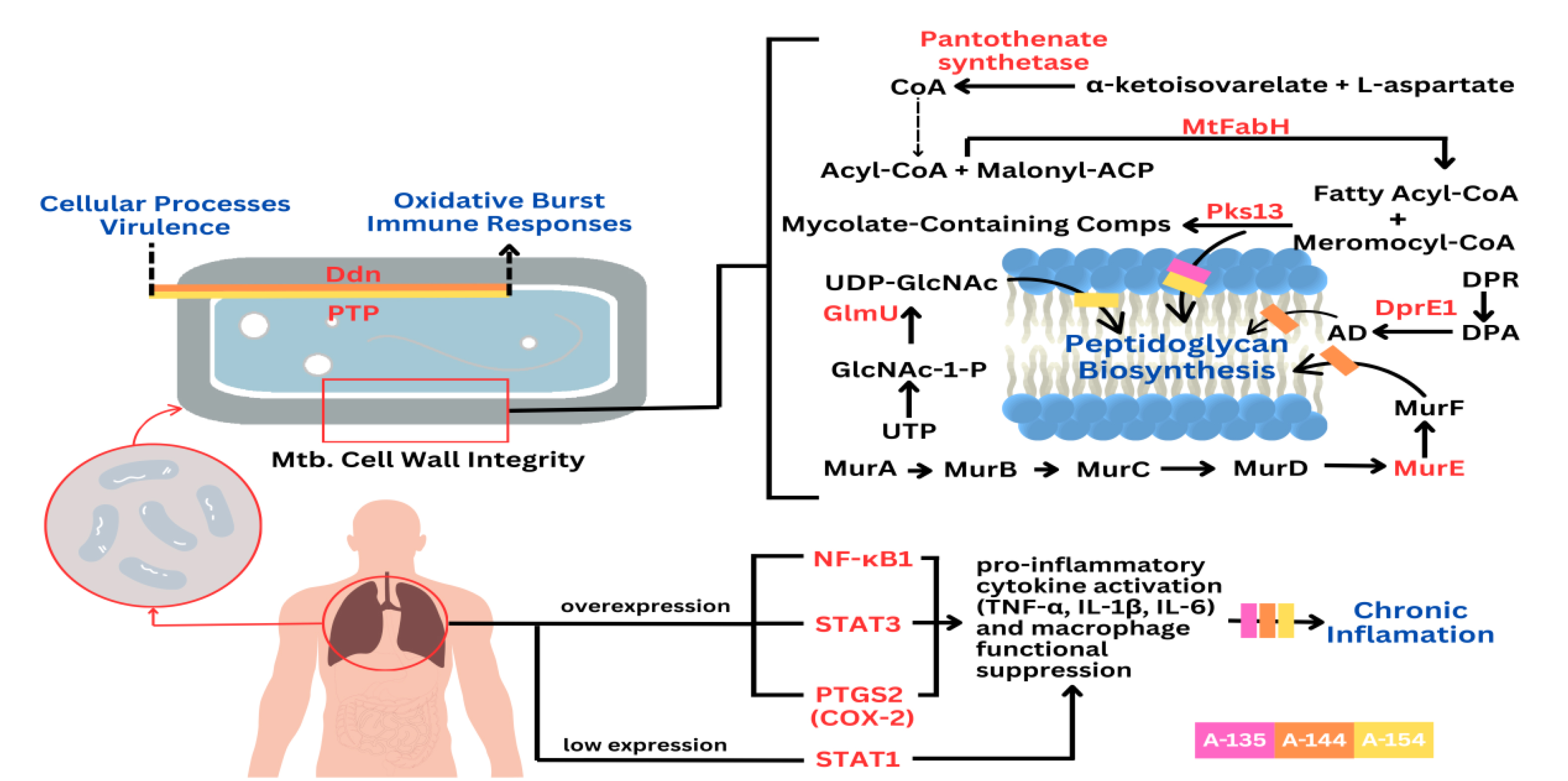
Proposed mechanism of action of combined halogenated benzylidene-cyclohexanone analogs a-135, a-144, and a-154 in counteracting Mycobacterium tuberculosis (Mtb) survivability and the chronic inflammatory manifestations of tuberculosis.
Nuclear Factor Kappa B Subunit 1 (NF-κB1), a key transcription factor in the NF-κB family, plays a pivotal role in host defense and the pathogenesis of TB. Upon Mycobacterium tuberculosis (Mtb) infection, alveolar macrophages identify pathogen-associated molecular patterns (PAMPs) and subsequently release pro-inflammatory cytokines, with TNF-α being a key example. The interaction of TNF-α with TNFR1 initiates a signaling pathway that includes IκB kinase (IKK). This enzyme then phosphorylates and degrades IκB, which allows NF-κB to move into the nucleus. Consequently, this process activates the transcription of genes that code for pro-inflammatory cytokines (such as IL-1β, IL-6, IL-12, and TNF-α), chemokines, and molecules that activate macrophages. Furthermore, NF-κB-mediated inflammation is crucial for granuloma formation, a defining characteristic of tuberculosis that limits the dissemination of Mtb, while also facilitating M1 macrophage polarization, Th1/Th17 responses, and inflammasome activation. Conversely, while excessive NF-κB activation can precipitate pathological inflammation and manifest as clinical symptoms of active tuberculosis, inadequate signaling undermines granuloma integrity and impairs bacterial control [50, 51]. Within this framework, the curcumin analogues A-135, A-144, and A-154, which were evaluated in this investigation, demonstrate inhibitory effects on NF-κB1, possibly by intervening in upstream processes, including IKK activation or IκB degradation. Consequently, these observations bolster their potential as adjunctive agents for tuberculosis therapy, achieved through the modulation of NF-κB-mediated immunopathology, without supplanting conventional antimicrobial interventions.
Signal Transducer and Activator of Transcription 3 (STAT3) serves as a pivotal transcription factor, significantly influencing immune responses in the context of Mycobacterium tuberculosis (Mtb) infection. This factor is activated by cytokines, including IL-10 and IL-6, which are released by immune cells during Mtb infection; following activation, STAT3 migrates to the nucleus, thereby regulating genes associated with cell proliferation, differentiation, and immune regulation. Within the framework of tuberculosis, STAT3 activation is observed in both infected macrophages and adjacent immune cells, thereby producing intricate immunomodulatory effects. Activated STAT3, in particular, downregulates the expression of crucial pro-inflammatory cytokines, such as IL-6, TNF-α, IFN-γ, and MIP-1β, while simultaneously inhibiting nitric oxide (NO) production through the repression of inducible nitric oxide synthase (iNOS). This suppression subsequently diminishes the microbicidal capabilities of macrophages, thereby promoting the intracellular survival of Mtb [52]. In this investigation, we assessed, through in silico analyses, a combination of compounds A-135, A-144, and A-154 that potentially impede STAT3 activation by disrupting its phosphorylation and nuclear translocation, with the aim of potentially reinstating pro-inflammatory mediator expression and augmenting macrophage antimicrobial functions. These results imply that targeting STAT3 with this compound combination could represent a promising adjuvant therapeutic approach in tuberculosis, contingent upon further experimental validation.
Signal Transducer and Activator of Transcription 1 (STAT1) is crucial for the host's defense against Mycobacterium tuberculosis (Mtb) through its mediation of interferon-gamma (IFN-γ) signaling. Following IFN-γ stimulation, STAT1 experiences phosphorylation, dimerization, and subsequent nuclear translocation, thereby activating interferon-stimulated genes (ISGs) via gamma-activated sequence (GAS) elements. This process subsequently triggers macrophage activation, nitric oxide production, and apoptosis within infected cells, thus restricting Mtb replication. Furthermore, STAT1 facilitates M1 macrophage polarization, which in turn enhances bactericidal activity. In the initial stages of tuberculosis (TB) infection, phosphorylated STAT1 initiates pro-apoptotic signaling; conversely, later stages are characterized by an accumulation of unphosphorylated STAT1, which hinders JAK1 phosphorylation and impairs apoptosis, thereby promoting immune evasion. Consequently, STAT1 deficiency is associated with heightened TB susceptibility, compromised granuloma formation, and increased bacterial loads.
Furthermore, molecules associated with STAT1 and regulatory RNAs have been identified as potential biomarkers for active tuberculosis (TB). Although this is significant, Mycobacterium tuberculosis (Mtb) can evade STAT1 responses by manipulating transcriptional coactivators, including CBP/p300, and selectively repressing genes responsive to IFN-γ [53, 54]. Notably, the combination of curcumin analogues A-135, A-144, and A-154, evaluated in this study, exhibits the potential to augment STAT1 activity, potentially through the promotion of its phosphorylation and nuclear translocation, thus counteracting Mtb's immune evasion tactics. While the exact molecular mechanism is yet to be fully defined, this combination may function by facilitating upstream kinase signaling or stabilizing STAT1–DNA binding. These observations imply a potential role for these compounds as adjunctive therapeutic agents, designed to strengthen host immune responses alongside standard anti-TB medications.
Cyclooxygenase-2 (COX-2), also known as prostaglandin-endoperoxide synthase 2, is a key enzyme in making prostanoids, especially prostaglandin E2 (PGE2). It significantly affects the immune response during Mycobacterium tuberculosis (Mtb) infection. In tuberculosis, COX-2 is increased in monocytes and macrophages after infection. This leads to more PGE2 being produced, which is a strong immunomodulator [55, 56]. The COX-2/PGE2 pathway has complex effects that depend on the situation. Early in the infection, PGE2 might helpcontrol excessive inflammation and encourage macrophage apoptosis. This could be protective by preventing excessive tissue damage and limiting bacterial spread.
Conversely, in the context of chronic or advanced tuberculosis (TB), increased levels of PGE2 can potentially hinder Th1 effector responses, diminish macrophage activation, and facilitate the proliferation of regulatory T cells (Tregs). This may compromise the host's capacity to manage Mycobacterium tuberculosis (Mtb) and, consequently, contribute to the advancement of the disease. Research has demonstrated that the inhibition of COX-2 through pharmacological agents, including indomethacin or etoricoxib, can decrease the population of Mtb-specific Tregs and influence cytokine production. However, this approach may also impede the macrophage's capacity to control mycobacterial infection, thereby implying that COX-2 activity is critical for optimal immune function under specific conditions. From a clinical perspective, dysregulated COX-2/PGE2 signaling is linked to both beneficial and adverse outcomes.At the same time, it can limit harmful inflammation and tissue destruction; excessive or prolonged activation may contribute to immune evasion by Mtb, increased bacterial burden, and more severe disease manifestations, such as persistent fever, weight loss, and impaired granuloma formation [55-57]. Our observations suggest that curcumin analogues A-135, A-144, and A-154 might reduce PTGS2 expression. This could be due to their interference with upstream NF-κB and MAPK signaling pathways. As a result, this interference could decrease PGE2 production and restore macrophage immunocompetence. Although the exact mechanism is not fully understood, this potential action highlights the usefulness of these compounds as additional treatments in host-directed TB therapy.
| Rank | Gene Symbol | Gene Name | Degree Score |
|---|---|---|---|
| 1 | EGFR | Epidermal Growth Factor Receptor | 90 |
| 2 | HSP90AA1 | Heat Shock Protein 90 Alpha Family Class A Member 1 | 86 |
| 3 | STAT3 | Signal Transducer and Activator of Transcription 3 | 84 |
| 4 | PTGS2 | Prostaglandin-Endoperoxide Synthase 2 | 71 |
| 5 | HSP90AB1 | Heat Shock Protein 90 Alpha Family Class B Member 1 | 69 |
| 6 | NF-κB1 | Nuclear Factor Kappa B Subunit 1 | 65 |
| 7 | EP300 | E1A Binding Protein P300 | 58 |
| 8 | APP | Amyloid Beta Precursor Protein | 51 |
| 9 | STAT1 | Signal Transducer And Activator Of Transcription 1 | 50 |
| 10 | NR3C1 | Nuclear Receptor Subfamily 3 Group C Member 1 | 45 |
Using a microwave-assisted method that followed green chemistry principles, we created new curcumin analogues. We then used an in silico approach to study their potential against tuberculosis (TB). By using molecular docking to target Mycobacterium tuberculosis (Mtb) proteins and network pharmacology to assess interactions with host-related targets, we found three promising candidates—A-135, A-144, and A-154. These showed good binding profiles and interacted with multiple targets related to TB development and symptoms. These compounds could be useful as additional treatments for TB. However, more research in the lab and in living organisms is needed to confirm their effectiveness and safety. Also, because curcumin and its derivatives often have formulation challenges, future work should focus on improving delivery systems to increase how well they work and how well they are absorbed (Table 3).
CONCLUSION
This investigation presents the effective microwave-assisted synthesis of halogenated benzylidene cyclohexanone curcumin analogs, specifically A-135 (yield: 81%), A-144 (yield: 58%), and A-154 (yield: 30%). These compounds demonstrated significant binding affinities toward essential Mycobacterium tuberculosis enzymes, thereby interfering with crucial pathways, including cell wall biosynthesis and lipid metabolism. Furthermore, network pharmacology analysis suggested their capacity to influence host immune responses through hub proteins NF-κB1, STAT1, STAT3, and PTGS2, thereby supporting a dual antimicrobial and host-directed mechanism. Despite its potential, this investigation is constrained by its computational methodology; although valuable for preliminary assessments, it lacks empirical confirmation within biological contexts. The identification of targets was dependent on existing databases and established Mycobacterium tuberculosis enzymes, potentially overlooking novel or poorly understood pathways. Furthermore, the network pharmacology analysis, despite its thoroughness, is predicated on predicted interactions, which may not accurately represent actual in vivo behavior. Additionally, while nine analogs were synthesized, only three were discussed in detail, leaving potential unexplored insights into the structure-activity relationship (SAR) of the remaining compounds. Future work should include experimental assays for target validation in cellular and animal models.
AUTHORS’ CONTRIBUTIONS
The authors confirm their contributions to the paper as follows: R.R.: Study conception and design; I.R., M.R.I., M.T.A.: Data collection; H.G.: Validation; P.M.: Analysis and interpretation of results; R.D.P.A.: Draft manuscript. All authors reviewed the results and approved the final version of the manuscript.
LIST OF ABBREVIATIONS
| TB | = Tuberculosis |
| MDR-TB | = multidrug-resistant TB GO = Gene Ontology |
| CRC | = Curcumin Research Center TLC = thin-layer chromatography IR = Infrared |
| NMR | = Nuclear Magnetic Resonance HRMS = High-Resolution Mass Spectrometry |
AVAILABILITY OF DATA AND MATERIALS
All data generated or analyzed during this study are included in this published article.
FUNDING
This work was supported by a grant from the Program of Post-Doctoral Batch II Research Grant 2022, Gadjah Mada University, Indonesia, Grant Number: 13602/UN1.P.II/Dit-Lit/PT.01.04/2022, 2 December 2022.
ACKNOWLEDGEMENTS
Declared none.


