All published articles of this journal are available on ScienceDirect.
Targeting WEE1 Kinase with Phytochemicals from Ampelocissus thyrsiflora (Blume) Planch: An Integrated In Silico and In Vitro Study Against Human Breast Cancer Cells
Abstract
Introduction
Breast cancer is the second most diagnosed cancer worldwide, with WEE1 kinase recognized as a key regulator of tumor progression. Natural products, particularly plant-derived metabolites, offer promising scaffolds for the development of novel WEE1 inhibitors.
Materials and Methods
Ethanol extracts of Ampelocissus thyrsiflora (Blume) Planch, a North Sumatran endemic plant, were obtained via maceration and subjected to phytochemical screening, PASS prediction, and Lipinski’s Rule of Five evaluation. Molecular docking and 100 ns molecular dynamics (MD) simulations were performed to investigate ligand–protein interactions. Cytotoxicity was assessed using the MTT assay on MCF-7 breast cancer cells.
Results
Phytochemical analysis confirmed the presence of alkaloids, flavonoids, saponins, tannins, glycosides, and steroids. Fourteen compounds exhibited strong predicted biological activity, and thirteen fulfilled drug-likeness criteria. Based on binding affinity and interactions with key WEE1 residues, four compounds were selected for MD simulations. Three demonstrated favorable MM-PBSA binding energies and stable interactions with critical residues. The ethanol extract showed an IC50 of 2011.25 ± 42.71 µg/mL against MCF-7 cells.
Discussion
The computational results highlight the potential of A. thyrsiflora metabolites as multi-target inhibitors of WEE1 kinase. Although the obtained IC50 value is comparatively high, the result provides an important preliminary basis for subsequent investigations and potential optimization of its active constituents.
Conclusion
Although the ethanol extract of A. thyrsiflora exhibited a high in vitro IC50, the in silico results provide a strong rationale for further exploration of its metabolites as potential WEE1-targeted anticancer agents.
1. INTRODUCTION
Cancer is a deadly disease that arises from genetic alterations in cells, leading to uncontrolled proliferation and tumour development [1]. According to GCO data, breast cancer is the second most frequently diagnosed malignancy, accounting for more than 11.6% of all cancers in women [2, 3]. Among the critical regulators of cell cycle progression is WEE1 protein kinase, which negatively regulates CDK1 and enforces the G2/M checkpoint, allowing cells time to repair DNA damage before mitosis. However, in cancer cells, this checkpoint is often exploited as a survival mechanism, enabling damaged and genetically unstable cells to persist [4, 5]. Targeting WEE1 has thus emerged as an urgent therapeutic strategy, as its inhibition can abrogate the G2 arrest, forcing DNA-damaged cells to prematurely enter mitosis and undergo apoptosis. This mechanism not only accelerates cell death but also sensitizes tumor cells to DNA-damaging chemotherapies and radiotherapy. The development of selective WEE1 inhibitors is therefore a high-priority area in cancer research, as they offer a more effective antitumor response compared to non-selective agents [6-8].
Adavosertib is a highly selective WEE1 inhibitor that increases tumor cell sensitivity to chemotherapeutic agents and replication stress [9, 10]. However, chemotherapy-based treatments are often associated with severe side effects such as nephrotoxicity, hepatotoxicity, and cardiotoxicity, and can also damage healthy DNA [11]. Consequently, there is a need for new, more targeted therapies with fewer side effects, such as those derived from natural metabolite compounds for the inhibition of cancer cell growth.
Ampelocissus thyrsiflora (Blume) Planch is a typical North Sumatran plant widely used in traditional medicine by the Karo people [12]. Phytochemical studies have shown that A. thyrsiflora contains various bioactive compounds, including alkaloids, flavonoids, saponins, tannins, and glycosides, which possess antioxidant and anticancer properties. Previous studies have also demonstrated its potent antioxidant activity and therapeutic benefits in tissue repair, reduction of lung inflammation, and hepatoprotection [13-15]. Additionally, toxicity screening using the Brine Shrimp Lethality Test (BSLT) revealed cytotoxicity, with an LC50 of 187.086 ppm against Artemia salina Leach, indicating that the plant contains compounds with bioactivity worth exploring [16].
Building on these findings, A. thyrsiflora has previously been reported to exhibit activity against SIRT1 and AMPK, which are associated with anticancer pathways [17]. Quercetin, a secondary metabolite contained in A. thyrsiflora, was known to suppress WEE1 expression in vitro, while isorhamnetin also effectively downregulated WEE1 expression [18, 19]. This suggests its potential as a WEE1 inhibitor. Molecular docking is a predictive, cost-effective method for identifying molecular interactions between plant-derived compounds and target proteins by evaluating binding affinity and stability. This technique facilitates the early-stage discovery of promising drug candidates [20]. Given this context, we conducted an experimental study integrating computational and in vitro assays to identify potential WEE1-targeting compounds from A. thyrsiflora leaves. These findings may contribute to new insights into the therapeutic potential of this underexplored plant species in breast cancer treatment.
2. MATERIALS AND METHODS
2.1. Materials
The materials used include a computer with ASUS ROG Zenith II Extreme Alpha, AMD Ryzen Threadripper 3970X, NVIDIA GeForce RTX 4090, AutoDock Vina, PyMOL, Biovia Discovery Studio [21], GROMACS, g-mmpbsa, PASS Online website (http://way2drug.com/Pass Online/), Lipinski's Rule of Five websites (https://www. scfbioiitd.res.in/software/drugdesign/lipinski.jsp), pkCSM Online Tools website (https://biosig.lab.uq.edu.au/pkcsm/), CHARMM-GUI (https://www.charmm-gui.org/), PROTOX-II website (https://tox.charite.de/protox3/), Protein Data Bank website (https://rcsb.org/), and PubChem website (https://pubchem.ncbi.nlm.nih.gov/).
2.2. Methods
This study employed an experimental, exploratory design to evaluate the interaction between phytochemicals from Ampelocissus thyrsiflora and the WEE1 kinase using in silico approaches and in vitro cytotoxicity assays. The study involved phytochemical characterization, molecular docking, molecular dynamics simulations, and MTT assay on MCF-7 cells. The primary variables assessed included binding energy, interaction residues, and IC50 values.
2.3. Extraction of A. thyrsiflora Leaves
Plant identification was carried out at the Badan Riset dan Inovasi Nasional (BRIN) under the reference number B-3283/II.6.2/IR.01.02/9/2024. The extraction process was carried out using 5 liters of 96% ethanol as the solvent for 500 grams of powdered A. thyrsiflora. The maceration was conducted at room temperature for 3 × 24 hours with stirring every 8 hours. The extract was filtered using filter paper to separate the filtrate from the residue. The filtrate was then concentrated using a rotary evaporator to obtain a thick extract. The yield of the extract was subsequently calculated [22, 23].
2.4. Phytochemical Screening of Ethanol Extract of A. thyrsiflora Leaves
Phytochemical screening of the ethanol extract of A. thyrsiflora leaves was conducted to detect the presence of secondary metabolites, including alkaloids, flavonoids, saponins, tannins, glycosides, and steroids. Alkaloids were identified using Mayer’s, Dragendorff’s, and Bouchardat’s reagents [24]. Flavonoids were detected by mixing the ethanol-dissolved extract with concentrated HCl and magnesium. Saponins were confirmed by the formation of stable foam. Tannins were identified upon the addition of 0.1% ferric chloride. Glycosides were detected by the treatment with NaOH [25], while steroids were confirmed through the Liebermann–Burchard test [26].
2.5. Chemical Compounds
Based on LC-HRMS analysis, 39 secondary metabolite compounds were identified in the ethanol extract of A. thyrsiflora, consisting of: Valproic Acid; Phenyl salicylate; Ribonic acid; Ethylparaben; 3',4'-dimethoxy-alpha-naphthoflavone; 6,6,9-trimethyl-3-(3-methyloctan-2-yl)-7,8,9,10 tetra hydrobenzo[c]chromen-1-ol;Dipropyleneglycol methyl ether acetate; Arbutin; 5-[6-hydroxy-5-(3-methylbut -2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol; Lauric acid; 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol; Norethindrone acetate; 2, 4, 5-trihydroxy-7, 8-dioxonaphthalen-1-olate; (2S, 3S)-2,3-Dihydro-2-[2-hydroxy-4-(betaDglucopyranosyloxy) phenyl]-3-(3, 5-dihydroxyphenyl) -4-[2-[4-(beta-D-glucopyranosy loxy) phenyl] ethenyl]benzofuran-6-ol; 3-hydroxy-3 methyl butanoate; Acetic acid; 1-Naphthylbis (9-anthryl) methylcation; 4-phenylethynylnaphthalic anhydride; [(2S,3R,4S,5S,6R) -3,4,5-trihydroxy-6-[[(2R,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxymethyl]oxan-2-yl] (E)-3-phenylprop-2-enoate; ethane; (13Z, 16Z)-docosa-13, 16-dienoic acid; Carthamoleusterone; 5-propan-2-yl-1, 3-benzodioxole; 3-[(2-carboxyacetyl) oxymethoxy]-3-oxopropanoic acid; Myricetin; Mearnsitrin; Quercetin; Kaempferol; 5,8-dihydroxy-2-(4-hydroxyphenyl)-7-methoxy-3-{[(2S,3R,4R, 5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}-4H-chromen-4-one; Isorhamnetin; Oleamide; Hexadecanamide; Stearamide; 1-Stearoylglycerol; Tridemorph; Erucamide; Bis(2-ethylhexyl) phthalate; Stearic acid; DL-α-Tocopherol.
2.6. Preparation of the Biology Activity Prediction
SMILES were first retrieved from PubChem and submitted to the PASS Online. Activating the prediction would show all the results [27].
2.7. Preparation of Physicochemical Prediction
All compounds were saved in *pdb format from PubChem and subjected to Lipinski's Rule of Five. On the web, the 3D structures in *pdb format should be submitted after the pH is set to 7 [28].
2.8. Molecular Docking
Docking was performed using AutoDock Vina with the 3D structure of the WEE1 receptor (PDB ID: 5VC5), while a total of 39 compounds from A. thyrsiflora obtained from PubChem were docked, and the docking process was conducted twice. The receptor was separated from the native ligands and water molecules, polar hydrogens were added, and Gasteiger charges were computed. Furthermore, a 30 × 30 × 30 Å grid box was used in the docking process [29, 30]. After obtaining the docking results, the lowest binding affinity was selected. Pymol software was used to examine the RMSD value. Visualization of the docking results was then performed using Biovia Discovery Studio [31]. Data were analyzed using IBM SPSS Statistics, and the mean ± standard deviation (SD) value.
2.9. Molecular Dynamics Simulations
Molecular dynamics (MD) simulations were generated from docking output and subsequently refined with CHARMM-GUI (https://www.charmm-gui.org/) to produce the topology files required for the MD simulations. The GROMACS force field was applied to define the systems and ligand parameters. The system temperature was maintained at 310°K throughout the simulation [32]. The simulations were executed using GROMACS 2022.4. An initial energy minimization was conducted to stabilize the system. The production phase involved a 100 ns simulation with a 4 fs time increment. Various metrics were analyzed, such as RMSD, RMSF, SASA, hydrogen bonding, and MMPBSA, and the results were visualized with GMXTools and g-mmpbsa [33, 34]. Data were analyzed using IBM SPSS Statistics, and the mean ± standard deviation (SD) value.
2.10. Ethics Approval
This study was conducted in accordance with the principles of the Declaration of Helsinki. Ethical approval was obtained from the Animal Research Ethics Committee of the Faculty of Mathematics and Natural Sciences, Universitas Sumatera Utara, Indonesia (Approval No. 0950/KEPH-FMIPA/2024). All experimental procedures were performed in compliance with institutional ethical guidelines.
2.11. Cytotoxicity Assay
MCF-7 cells were obtained from the Laboratory of Parasitology, Faculty of Medicine, Public Health, and Nursing, Universitas Gadjah Mada, Indonesia. The cells were seeded in 96-well plates at a density of 1 × 104 cells/100 µL and incubated for 24 hours in a CO2 incubator at 37 °C. The ethanol extract of A. thyrsiflora leaves and adavosertib, dissolved in DMSO, were then added to the medium at concentrations of 1000, 500, 250, 125, and 62.5 µg/mL, with three replicates per concentration. After 24 hours of incubation, the medium was removed, and 100 µL of 0.5 mg/mL MTT solution was added to each well. After 6 hours, 10% SDS was added. The optical density (OD) was measured using a microplate reader at 595 nm [35, 36].
3. RESULTS AND DISCUSSION
3.1. Ethanol Extract of A. thyrsiflora Leaves
The extraction of A. thyrsiflora was carried out using 96% ethanol. This choice was based on the characteristics of 96% ethanol, which can extract both polar and non-polar compounds, as well as the advantages of the maceration method, which is easy to perform and does not require heating, thus preventing the degradation of the compounds present in the sample [37]. The extraction yielded a concentrated extract of 95.91 grams [38]. The extract yield was then calculated by comparing the final weight (weight of the concentrated extract) to the initial weight (weight of the A. thyrsiflorapowder used), and multiplying by 100% [39]. Therefore, the extract yield obtained was 19.1%. The ethanol extract yield met the requirements, as the standard for a concentrated extract yield is greater than 10%. Yield represents the percentage of the material remaining after extraction and serves to assess the effectiveness of the extraction [40].
3.2. Phytochemical Screening of A. thyrsiflora Leaves
Phytochemical screening was conducted for alkaloids, flavonoids, saponins, tannins, glycosides, and steroids using appropriate reagents. The results are shown in Table 1.
Table 1.
| Sample | Compounds | Results |
|---|---|---|
| Ethanol extract of A. thyrsiflora | Alkaloids | + |
| Flavonoids | + | |
| Saponins | + | |
| Tannins | + | |
| Glycosides | + | |
| Steroids | + |
As shown in Table 1, all phytochemical tests were positive for the six secondary metabolites. These positive results were indicated by changes in color, the formation of precipitates, and the development of foam. This is in line with the phytochemical screening of A. thyrsiflora leaves conducted by Marbun [41], who reported positive results for all secondary metabolites.
3.3. Biology Activity Prediction
The prediction of biological activity was performed on the PASS Online website, which displays the probability of activity (Pa) for each pharmacological effect that could occur in the compounds, as shown in Table 2.
PASS Online is a database that predicts the activity spectrum of substances, which is used to predict the biological spectrum of compounds. If a compound has a bioactivity test value of Pa > 0.7, it has very high biological activity; if Pa < 0.5, the bioactivity is very low; and if 0.5 < Pa < 0.7, the bioactivity is moderate [42, 43]. According to that, 14 compounds fulfilled the requirement. Compounds that have good antineoplastic activity are 3',4'-dimethoxy-alpha-naphthoflavone; arbutin; 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene -1,3-diol; 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol; Myricetin; Mearnsitrin; Quercetin; Kaempferol. Compounds that have good Anticarcinogenic activity are (2S, 3S)-2, 3-Dihydro-2-[2-hydroxy-4-(beta-D-glucopyranosyloxy) phenyl]-3-(3, 5-dihydroxyphenyl) -4-[2-[4-(beta-D-glucopyranosyloxy) phenyl]ethenyl]benzofuran-6-ol; Myricetin; Mearnsitrin; Quercetin; Kaempferol; Isorhamnetin. In addition to the Ampelocissus thyrsiflora, some compounds have high antioxidant activity including 2S, 3S)-2, 3-Dihydro-2-[2-hydroxy-4-(beta-D-glucopyranosyloxy) phenyl]-3-(3, 5-dihydroxyphenyl) -4-[2-[4-(beta-D-glucopyranosyloxy) phenyl]ethenyl]benzofuran-6-ol; Myricetin; Mearnsitrin; Quercetin; Kaempferol; Isorhamnetin; DL-α-Tocopherol and apoptosis agonist are 3',4'-dimethoxy-alpha-naphthoflavone; 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol; 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol; Myricetin; Quercetin; Kaempferol; α-Tocopherol. Apoptosis is essential for cell development and for maintaining the stable internal state (homeostasis) of tissues, including morphological changes [44]. Biological activities in the form of antineoplastic, apoptosis agonist, antioxidant, and anticarcinogen possessed by compounds in A. thyrsiflora Leaves indicate that this plant has the potential to be utilized in the treatment of breast cancer [45].
3.4. Physicochemical Prediction
The rule of five (ROF) is a parameter used to evaluate drug-likeness or to predict the potential pharmacological or biological activity of a chemical compound with oral activity as a medicine [46], as shown in Table 3.
The description of + stands for requirements fulfilled, while – means requirements not fulfilled, physicochemical prediction results by Lipinski's Rule of Five of A. thyrsiflora. As has been written on Table 1, 13 compounds that fulfilled Lipinski's Rule of Five are valproic acid; phenyl salicylate; ethylparaben; 3',4'-dimethoxy-alpha-naphthoflavone; dipropyleneglycol methyl ether acetate; 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol; lauric acid; norethindrone acetate; 2, 4, 5-trihydroxy-7, 8-dioxonaphthalen-1 olate; kaempferol; isorhamnetin; hexadecanamide; 1-stearoylglycerol. A chemical compound is said to have good absorption or permeation if the Molar mass <500, the Number of H-bond acceptors <10, the Number of H-bond donors < 5, and Log P <5 [47]. If the compound has a molecular weight greater than 500 g/mol, it will be difficult to absorb and have low permeability [48].
| No | Compound | Pa | Biological Activity |
|---|---|---|---|
| 1 | 3',4'-dimethoxy-alpha-naphthoflavone | 0,734 | Antineoplastic |
| 0,743 | Apoptosis agonist | ||
| 2 | Arbutin | 0,829 | Anti-carcinogenic |
| 0,782 | Antioxidant | ||
| 0,787 | Antineoplastic | ||
| 3 | 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol | 0,718 | Antineoplastic |
| 0,835 | Apoptosis agonist | ||
| 4 | 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol | 0,746 | Antineoplastic |
| 0,795 | Apoptosis agonist | ||
| 5 | (2S, 3S)-2, 3-Dihydro-2-[2-hydroxy-4-(beta-D-glucopyranosyloxy) | 0,881 | Anti-carcinogenic |
| 0,723 | Antioxidant | ||
| 0,781 | Antineoplastic | ||
| 6 | [(2S,3R,4S,5S,6R)-3,4,5-trihydroxy-6-[[(2R,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxymethyl]oxan-2-yl] (E)-3-phenylprop-2-enoate | 0,865 | Anti-carcinogenic |
| 0,736 | Antioxidant | ||
| 0,861 | Antineoplastic | ||
| 7 | Carthamoleusterone | 0,781 | Antineoplastic |
| 8 | Myricetin | 0,784 | Anti-carcinogenic |
| 0,924 | Antioxidant | ||
| 0,841 | Antineoplastic | ||
| 0,915 | Apoptosis agonist | ||
| 9 | Mearnsitrin | 0,952 | Anti-carcinogenic |
| 0,898 | Antioxidant | ||
| 0,878 | Antineoplastic | ||
| 0,824 | Apoptosis agonist | ||
| 10 | Quercetin | 0,757 | Anti-carcinogenic |
| 0,872 | Antioxidant | ||
| 0,797 | Antineoplastic | ||
| 0,887 | Apoptosis agonist | ||
| 11 | Kaempferol | 0,715 | Anti-carcinogenic |
| 0,856 | Antioxidant | ||
| 0,791 | Antineoplastic | ||
| 0,881 | Apoptosis agonist | ||
| 12 | 5,8-dihydroxy-2-(4-hydroxyphenyl)-7-methoxy-3-{[(2S,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}-4H-chrome-4-one | 0,929 | Anti-carcinogenic |
| 0,888 | Antioxidant | ||
| 0,854 | Antineoplastic | ||
| 0,803 | Apoptosis agonist | ||
| 13 | Isorhamnetin | 0,779 | Anti-carcinogenic |
| 0,809 | Antioxidant | ||
| 0,803 | Antineoplastic | ||
| 0,88 | Apoptosis agonist | ||
| 14 | DL-α-Tocopherol | 0,967 | Antioxidant |
| 0,736 | Apoptosis agonist |
| No | Compound | MW | Log P | HBD | HBA | Molar Refractivity | Desc. |
|---|---|---|---|---|---|---|---|
| 1 | Valproic Acid | 144 | 2.28 | 1 | 2 | 40.94 | + |
| 2 | Phenyl salicylate | 214 | 2.61 | 1 | 3 | 59.5 | + |
| 3 | Ribonic acid | 166 | -2.85 | 5 | 6 | 32.74 | - |
| 4 | Ethylparaben | 166 | 1.56 | 1 | 3 | 44.06 | + |
| 5 | 3',4'-dimethoxy-alpha-naphthoflavone | 332 | 4.47 | 0 | 4 | 96.42 | + |
| 6 | 6,6,9-trimethyl-3-(3-methyl octan-2-yl)-7,8,9,10-tetrahydrobenzo[c]chromen-1-ol | 370 | 7.45 | 1 | 2 | 114.46 | - |
| 7 | Dipropylene glycol methyl ether acetate | 190 | 0.98 | 0 | 4 | 48.51 | + |
| 8 | Arbutin | 272 | -1.42 | 5 | 7 | 62.5 | - |
| 9 | 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol | 310 | 4.53 | 3 | 4 | 89.11 | + |
| 10 | Lauric acid | 200 | 3.99 | 1 | 2 | 59.47 | + |
| 11 | 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol | 410 | 8 | 1 | 1 | 128.23 | - |
| 12 | Norethindrone acetate | 340 | 4.06 | 0 | 3 | 95.42 | + |
| 13 | 2, 4, 5-trihydroxy-7, 8-dioxonaphthalen-1-olate | 221 | 0 | 3 | 6 | 49.61 | + |
| 14 | (2S, 3S)-2, 3-Dihydro-2-[2-hydroxy-4-(beta-D-glucopyranosyloxy) phenyl]-3-(3, 5-dihydroxyphenyl) -4-[2-[4-(beta-D-glucopyranosyloxy) phenyl]ethenyl]benzofuran-6-ol | 752 | -3.46 | 0 | 17 | 162.78 | - |
| 15 | 3-hydroxy-3 methyl butanoate | 118 | 0.23 | 2 | 3 | 28.55 | - |
| 16 | Acetic acid | 60 | 0.09 | 1 | 2 | 13.3 | - |
| 17 | 1-Naphthylbis (9-anthryl) methylation | 493 | 6.44 | 0 | 0 | 150.04 | - |
| 18 | 4-phenylethynylnaphthalic anhydride | 526 | 7.78 | 0 | 3 | 162.48 | - |
| 19 | [(2S,3R,4S,5S,6R)-3,4,5-trihydroxy-6-[[(2R,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxymethyl]oxan-2-yl] (E)-3-phenylprop-2-enoate | 456 | -2.1 | 6 | 11 | 106.55 | - |
| 20 | ethane | 30 | 1.02 | 0 | 0 | 11.34 | - |
| 21 | (13Z, 16Z)-docosa-13, 16-dienoic acid | 336 | 7.44 | 1 | 2 | 105.46 | - |
| 22 | Carthamoleusterone | 508 | 3.99 | 4 | 8 | 136.32 | - |
| 23 | 5-propan-2-yl-1, 3-benzodioxole | 696 | 8.31 | 0 | 7 | 192.14 | - |
| 24 | 3-[(2-carboxyacetyl) oxymethoxy]-3-oxopropanoic acid | 220 | -1.02 | 2 | 8 | 41.55 | - |
| 25 | Myricetin | 318 | 1.71 | 6 | 8 | 75.71 | - |
| 26 | Mearnsitrin | 478 | 0.3 | 7 | 12 | 111.41 | - |
| 27 | Quercetin | 302 | 2.01 | 5 | 7 | 74.05 | - |
| 28 | Kaempferol | 286 | 2.3 | 4 | 6 | 72.38 | + |
| 29 | 5,8-dihydroxy-2-(4-hydroxyphenyl)-7-methoxy-3-{[(2S,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}-4H-chrome-4-one | 440 | -1.32 | 0 | 11 | 92.07 | - |
| 30 | Isorhamnetin | 316 | 2.31 | 4 | 7 | 78.93 | + |
| 31 | Oleamide | 281 | 5.5 | 2 | 2 | 88.64 | - |
| 32 | Hexadecanamide | 255 | 4.95 | 2 | 2 | 79.5 | + |
| 33 | Stearamide | 283 | 5.73 | 2 | 2 | 88.74 | - |
| 34 | 1-Stearoylglycerol | 358 | 4.5 | 3 | 4 | 103.65 | + |
| 35 | Tridemorph | 297 | 5.4 | 0 | 2 | 92.9 | - |
| 36 | Erucamide | 337 | 7.06 | 2 | 2 | 107.11 | - |
| 37 | Bis(2-ethylhexyl) phthalate | 390 | 6.2 | 0 | 4 | 112.68 | - |
| 38 | Stearic acid | 337 | 7.06 | 2 | 2 | 107.11 | - |
| 39 | DL-α-Tocopherol | 430 | 8.84 | 1 | 2 | 134.39 | - |
Structure of WEE1 after receptor preparation.
3.5. Receptor Preparation and Docking Validation
Protein used for this study is WEE1 (PDB ID: 5VC5), homo sapiens organism, resolution value 1.93 Å, which can be seen in Fig. (1).
3.6. Docking Validation
The RMSD (Root Mean Square Deviation) value for WEE1 was 1.195 Å, indicating that the docking results were valid, as the RMSD value was below the accepted threshold of 2.0 Å [49] (Fig. 2). The prepared receptors were then docked with ligands using AutoDock Vina to obtain amino acid residues, binding affinities, and standard deviations.
3.7. Docking Result with Biovia Discovery Studio
Biovia Discovery Studio visualized the biomolecular interactions, allowing researchers to analyze the interactions and bonds created between ligand and protein in the binding pockets. Table 4 reports the mean binding affinity as the average of two binding affinity measurements per compound, since each was conducted twice.
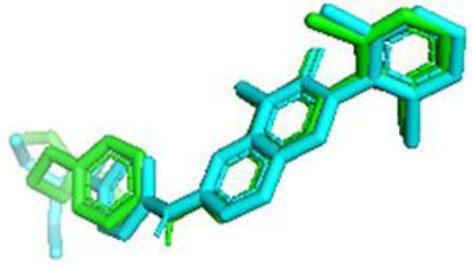
The validation result of the native ligand (green) and ligand docking (blue) of WEE1.
| No | Compounds | H-Bond Interaction | Hydrophobic Bond Interaction | Binding Affinity |
|---|---|---|---|---|
| 1 | Adavosertib | PHE310, GLU377, ASP463 | ILE305, VAL313, ALA326, LYS328, VAL360, PHE433 | -10.09 ± 0.38 |
| 2 | Native ligand | GLU377, CYS379 | ILE305, VAL313, ALA326, LYS328, ASP386 | -9.54 ± 0.24 |
| 3 | Valproic Acid | LYS331 | PHE310 | -3.48 ± 0.28 |
| 4 | Phenyl salicylate | - | ALA326, LYS328, ILE374, CYS379, PHE433 | -7.65 ± 0.07 |
| 5 | Ribonic acid | MET425, ASP426, LEU483 | - | -5.73 ± 0.1 |
| 6 | Ethylparaben | - | ALA326, VAL360, CYS379, PHE433 | -6.11 ± 0.01 |
| 7 | 3',4'-dimethoxy-alpha-naphthoflavone | GLY306, SER307, ASN376 | ILE305, VAL313, ALA326 | -9.03 ± 0.0 |
| 8 | 6,6,9-trimethyl-3-(3-methyloctan-2-yl)-7,8,9,10-tetrahydrobenzo[c]chromen-1-ol | SER307, GLU309 | GLY308, PHE310 | -2.53 ± 0.57 |
| 9 | Dipropylene glycol methyl ether acetate | PHE310, LYS328, LYS331 | - | -4.09 ± 0.09 |
| 10 | Arbutin | SER307, SER312, ARG329 | - | -4.96 ± 0.15 |
| 11 | 5-[6-hydroxy-5-(3-methylbut-2-enyl)-1-benzofuran-2-yl] benzene-1,3-diol | PRO333 | PHE310, LYS331, ASP339. HIS466 | -7.22 ± 0.0 |
| 12 | Lauric acid | LYS328 | ILE305, VAL313, ALA326, VAL360, CYS 379, PHE433 | -5.77 ± 0.09 |
| 13 | 4,4-Dimethyl-5 alpha-cholesta-8,14,24-trien-3 beta-ol | - | PHE314 | -6.25 ± 0.48 |
| 14 | Norethindrone acetate | - | PHE310, LYS331 | -6.08 ± 0.26 |
| 15 | 2, 4, 5-trihydroxy-7, 8-dioxonaphthalen-1-olate | PHE310, LYS331 | GLU309, ASP339 | -6.39 ± 0.58 |
| 16 | (2S, 3S)-2, 3-Dihydro-2-[2-hydroxy-4-(beta-D-glucopyranosyloxy) phenyl]-3-(3, 5-dihydroxyphenyl) -4-[2-[4-(beta-D-glucopyranosyloxy) phenyl]ethenyl]benzofuran-6-ol | LYS304, SER307, GLU309, SER312, ARG329 | GLY308, PHE310 | -6.89 ± 0.16 |
| 17 | 3-hydroxy-3-methyl butanoate | SER307, GLY308, GLY311 | - | -1.94 ± 0.18 |
| 18 | Acetic acid | GLU309 | - | -0.98 ± 0.0 |
| 19 | 1-Naphthylbis (9-anthryl) methylcation | - | VAL338 | -2.77 ± 0.05 |
| 20 | 4-phenylethynylnaphthalic anhydride | - | VAL338 | -2.58 ± 0.02 |
| 21 | [(2S,3R,4S,5S,6R)-3,4,5-trihydroxy-6-[[(2R,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxymethyl]oxan-2-yl] (E)-3-phenylprop-2-enoate | GLY308, GLU309, LYS328, ASP463 | PHE310, VAL313, PRO333, ASP339 | -6.39 ± 0.58 |
| 22 | Ethane | - | PHE310 | -1.41 ± 0.0 |
| 23 | (13Z, 16Z)-docosa-13, 16-dienoic acid | SER312, ARG329 | - | -3.72 ± 0.0 |
| 24 | Carthamoleusterone | PHE310 | - | -1.68 ± 0.39 |
| 25 | 5-propan-2-yl-1, 3-benzodioxole | - | ARG329 | -3.88 ± 0.0 |
| 26 | 3-[(2-carboxyacetyl) oxymethoxy]-3-oxopropanoic acid | LYS304, GLY311, SER312, ARG329, HIS371 | - | -4.45 ± 0.1 |
| 27 | Myricetin | PHE310, ALA343 | ASP339 | -6.46 ± 0.0 |
| 28 | Mearnsitrin | LYS331 | LYS304 | -5.94 ± 0.61 |
| 29 | Quercetin | ASP 339 | GLU309, PHE310, ASP339 | -6.7 ± 0.0 |
| 30 | Kaempferol | PHE310, LYS328, LYS331 | PRO333, ASP339 | -6.55 ± 0.23 |
| 31 | 5,8-dihydroxy-2-(4-hydroxyphenyl)-7-methoxy-3-{[(2S,3R,4R,5R,6S)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}-4H-chromen-4-one | PHE310, LYS328, PRO333, LEU334, SER337, ASP339 | GLU309 | -7.13 ± 0.19 |
| 32 | Isorhamnetin | PHE310, ASP426, LYS428 | ASP339, HIS466 | -6.46 ± 0.01 |
| 33 | Oleamide | SER312, ARG329 | PHE310 | 0.8 ± 1.63 |
| 34 | Hexadecanamide | SER307, SER312 | PHE310 | -2.81 ± 0.23 |
| 35 | Stearamide | SER304, SER312 | PHE314 | -1.69 ± 0.15 |
| 36 | 1-Stearoylglycerol | GLY308, GLY311 | PHE314 | -2.97 ± 0.06 |
| 37 | Tridemorph | GLY308 | PHE310, PHE314 | -2.38 ± 0.24 |
| 38 | Erucamide | ASP339 | VAL338 | -2.17 ± 0.03 |
| 39 | Bis(2-ethylhexyl) phthalate | - | PHE310, VAL338, ASP339 | -2.03 ± 0.09 |
| 40 | Stearic acid | ASP339 | VAL338 | -1.64 ± 0.14 |
| 41 | DL-α-Tocopherol | - | PHE310 | -3.84 ± 0.65 |
Binding affinity is a value that indicates the accuracy of the bond between the ligand and the target protein [50]. The lower the binding affinity, the stronger the bond between the ligand and the receptor, making it a promising therapeutic candidate [51]. The binding affinity of the WEE1 protein ranged from 0.8 to -10.09. The standard drug, Adavosertib, and native ligand (96M) exhibit the lowest binding affinity.
Based on previous research, the binding modes between WEE1 and WEE1 inhibitors include the pyrimidine nitrogen adjacent to an amino group forming H-bonds with the CYS379 backbone, hydroxyl groups on the pyridine side chain forming H-bonds with ASP463, and the pyrazolopyridinone core establishing π-stacking interactions with PHE433 [52]. Other hydrogen bonds that contribute to inhibitory activity have been reported with ILE305, LYS328, and ASP339 [53-55]. In our docking results, several compounds interacted with the same critical amino acid residues within the WEE1 binding pocket. Among them, four compounds: kaempferol, myricetin, quercetin, and isorhamnetin were prioritized not only because they displayed relatively favorable binding affinities, but also because they consistently engaged key residues known to be essential for WEE1 inhibition [56]. Importantly, affinity does not necessarily predict activity, as binding ligands may behave as either agonists or competitive inhibitors depending on their interactions within the active site. Therefore, the quality and specificity of amino acid interactions play a crucial role in determining whether a ligand truly disrupts the biological function of the target protein [57]. This dual criterion of binding strength and residue-specific interactions provided a strong rationale for selecting these four flavonoids for subsequent molecular dynamics simulations over 100 ns to further evaluate their stability and binding modes.
3.8. Molecular Dynamics Simulations
Molecular dynamics simulations were done using GROMACS and CHARMM-GUI for input file preparation. MD simulations were performed for a 100 ns trajectory. The visualization of the ligand throughout the 100 ns simulations can be seen in Fig. (3).
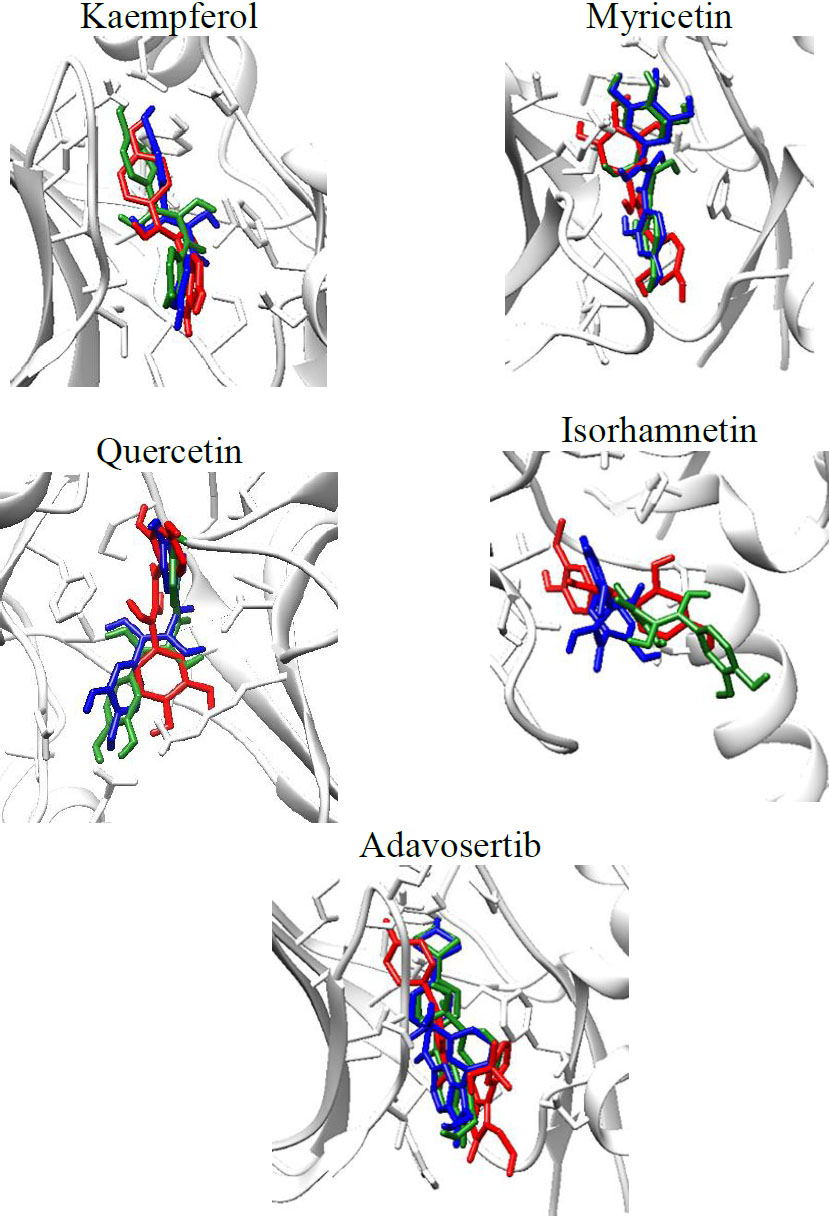
Visualization of the WEE1-ligand complexes throughout the 100 ns simulations, red (0 ns), green (50 ns), blue (100 ns).
From the visualization above, it can be seen that the ligand stays in the binding site throughout the 100 ns simulation. Analysis of the RMSD, RMSF, SASA, and Hydrogen bonds formed during the simulations was also done. The results are shown in Fig. (4).
RMSD is an atom's average movement or displacement at a specific time interval that provides data on the average distance between ligands and proteins. Based on the results, the RMSD of the WEE1-ligand complex shows a slight increase at 0.5 ns, followed by greater stability thereafter. The interaction between ligand-receptor is considered stable if the RMSD value is under 2.5 nm, in which these results show a stable interaction [58, 59].
Root-mean-square fluctuation (RMSF) aims to evaluate the mobility of molecular systems. The RMSF value measures the degree of structural flexibility or rigidity of individual atoms within a molecule or system [60]. The RMSF data show that the fluctuations of the WEE1-ligands happen between 0.03 and 0.5 nm, with the activation segment peaking in 370 to 410 residues. The fluctuation in the activation segment indicates the ability to bind ligands and elicit a biological response; these results align with findings that the WEE1 binding pocket involves ASN 376, GLU 377, CYS 379, and ASP 386 [52]. A high RMSF value indicates a flexible region, while a low RMSF value indicates the rigidity of the amino acid [61, 62].
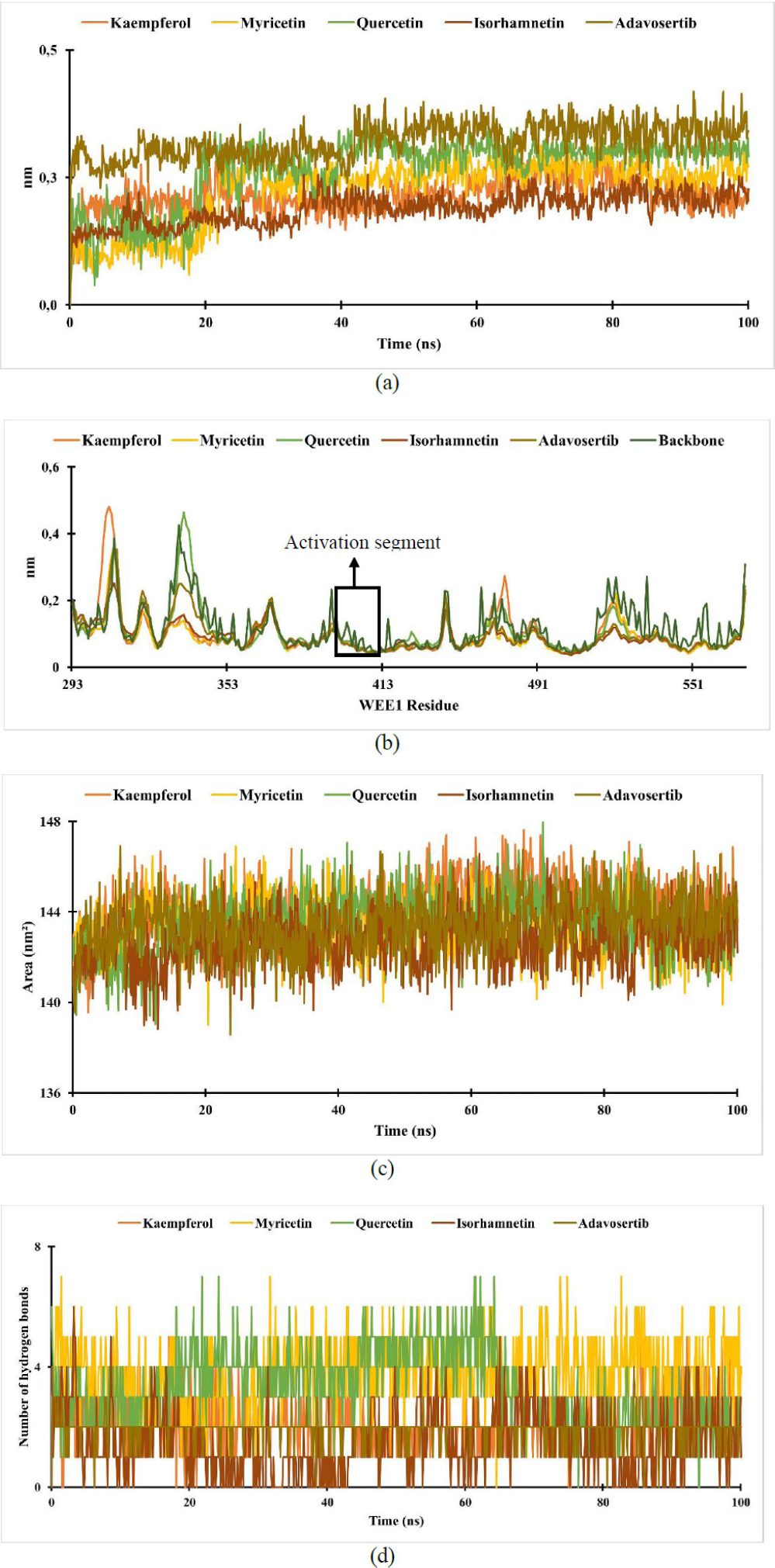
Analysis of the (a) RMSD of WEE1-ligand complex; (b) RMSF of WEE1-ligand complex; (c) SASA of WEE1-ligand complex; (d) hydrogen bond of WEE1-ligand complex.
The solvent-accessible surface area (SASA) is the surface area of a molecule that can come into contact with the solvent [63]. From the results, the SASA of the ligand-receptor complexes remained stable throughout the simulations, ranging from 138 to 148 nm2. Higher SASA values indicate larger, looser proteins and more flexible protein conformations, which correlate with increased interaction potential with other molecules, such as ligands [64].
Hydrogen bonding is an interaction that occurs in hydrogen atoms contained in a pair of other atoms that have an affinity for electrons [65]. The ligand will remain close to the protein-binding site through permanent hydrogen bonds [61]. In the results, the number of hydrogen bonds formed during the simulation ranged from 0 to 7 for WEE-ligand complexes.
Further analysis was done to obtain each complex's amino acid residues and MMPBSA value. The results are shown in Table 5.
MMPBSA analysis was then done to estimate ligand-binding affinities. Evdw (Van der Waals energy) accounts for the weak intermolecular forces and significantly contributes to the overall binding free energy. Electrostatic energy (Eele) plays a crucial role initially but often cancels out the polar solvation energy due to the solvent's screening effect. Epolar (Polarsolvation energy) represents the electrostatic interactions between the solute and the surrounding solvent. Enon-polar (Nonpolar solvation energy) encompasses the effects of creating a cavity in the solvent and the subsequent solvent rearrangement around the solute [66-68]. The total energy is summed from the components to produce each component's average total free energy. So, to calculate the binding free energy, the total free energy of the receptor and ligand is subtracted from the total free energy of the complex [69]. The results demonstrated that kaempferol, myricetin, and quercetin exhibited lower binding free energies than the standard drug adavosertib, indicating stronger binding affinities for the WEE1 receptor. These flavonoids are widely recognized for their anticancer activities, with numerous reports describing their ability to induce apoptosis, inhibit proliferation, and modulate signaling pathways in breast cancer cells. Notably, quercetin has previously been reported to inhibit WEE1 activity, supporting the credibility of our computational predictions [17]. Both quercetin and the other flavonoids formed key interactions with critical WEE1 amino acid residues, including hydrogen bonds with GLU377 and CYS379, as well as hydrophobic interactions with ALA326 and CYS379, which are known to contribute to WEE1 inhibition [52]. To validate these computational findings, an in vitro cytotoxicity assay using the MTT method was conducted on the MCF-7 breast cancer cell line. The combined in silico and in vitro results suggest that the metabolite compounds present in A. thyrsiflora leaves have promising potential as WEE1 inhibitors, warranting further investigation.
3.9. Cytotoxicity Assay
The cytotoxicity result of the ethanol extract of A. thyrsiflora leaves against MCF-7 cell lines can be seen in Fig. (5) and Table 6.
| Compounds | Amino Acid Residue | MMPBSA | |||||
|---|---|---|---|---|---|---|---|
| H-bond Interaction | Hydrophobic Bonds Interaction | ΔEVDW | ΔEELE | ΔEpolar | ΔEnon polar | ΔEGTOTAL | |
| Kaempferol | LYS328, ASN376, GLU377, CYS379 | VAL313, ALA326, CYS379 | -36.90 ± 0.42 | -35.16 ± 0.95 | 46.88 ± 0.96 | -4.93 ± 1.00 | -30.11 ± 0.57 |
| Myricetin | GLU377, CYS379 | ALA326, CYS379 | -38.31 ± 0.57 | -48.41 ± 0.55 | 59.58 ± 0.51 | -5.79 ± 0.09 | -32.93 ± 0.71 |
| Isorhamnetin | GLY308, ASP386 | ILE305 | -25.79 ± 0.31 | -21.33 ± 0.08 | 36.98 ± 0.14 | -4.13 ± 0.06 | -14.27 ± 0.30 |
| Quercetin | GLU346, GLU377, CYS379, GLY382 | ILE305, VAL313, ALA326, LYS328, PHE433 | -36.11 ± 0.04 | -36.43 ± 0.31 | 49.94 ± 0.07 | -5.35 ± 0.08 | -27.95 ± 0.43 |
| Adavosertib | ASN376, GLU377, TYR378, CYS379 | ILE305, VAL313, ALA326, VAL360, CYS379, PHE433 | -45.85 ± 0.07 | -29.18 ± 0.05 | 59.14 ± 0.07 | -6.43 ± 0.33 | -22.31 ± 0.27 |
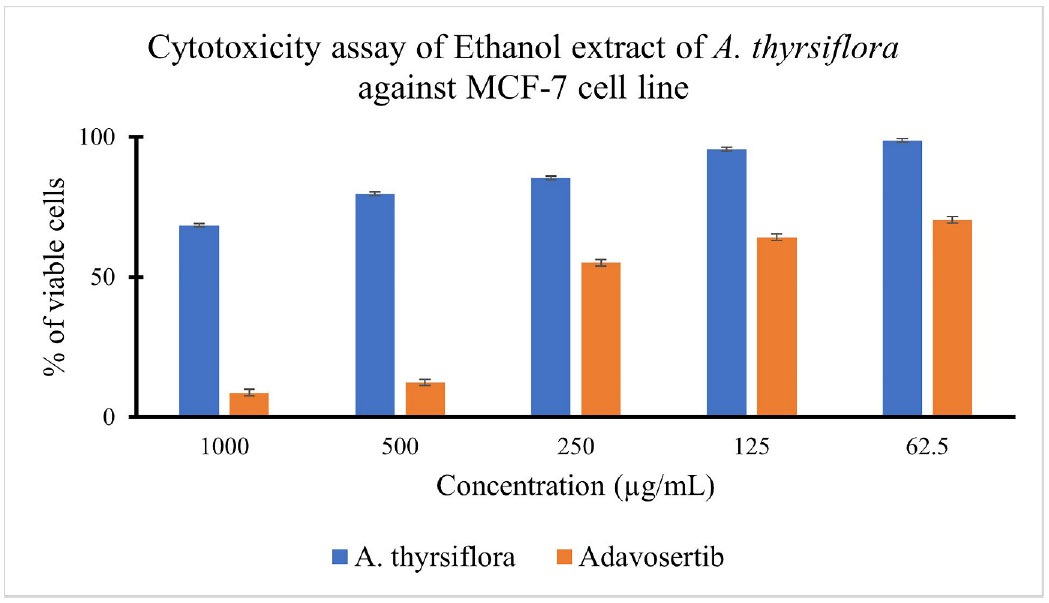
% of viable cells of ethanol extract of A. thyrsiflora and Adavosertib treatment against the MCF-7 cell line.
| Value | Ethanol Extract of A. thyrsiflora | Adavosertib |
|---|---|---|
| IC50 (µg/ml) | 2011.25 ± 42.71 | 175.98 ± 1.66 |
The MTT assay is based on the ability of mitochondrial dehydrogenases, primarily succinate dehydrogenase in metabolically active cells, to reduce the yellow tetrazolium salt, MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide), into insoluble purple formazan crystals. Since only viable cells with intact mitochondria and cell membranes can perform this reduction, the amount of formazan formed is directly proportional to the number of living cells and can be quantified using a spectrophotometer [70, 71]. In addition to determining cytotoxic effects, the MTT assay provides the half-maximal inhibitory concentration (IC50), which is a standard parameter used to evaluate the potency of cytotoxic agents in cell-based assays [72]. As shown above, cell viability decreased with increasing extract concentration. At 1000 µg/mL of the ethanol extract of A. thyrsiflora, cell viability dropped to 68%, whereas treatment with 1000 µg/mL of Adavosertib reduced cell viability by 91%. Adavosertib is a well-known WEE1 inhibitor with an IC50 of 175.98 ± 1.66 µg/mL, and its activity is associated with disruption of the G2/M checkpoint, inhibition of DNA damage repair, and induction of apoptosis [73]. The calculated IC50 value indicated that the extract exhibited limited cytotoxic activity, according to the classification criteria set by the U.S. National Cancer Institute [74]. Although the ethanol extract of A. thyrsiflora demonstrated a high IC50 value, the observed dose-dependent decrease in MCF-7 cell viability suggests the presence of bioactive constituents with potential anticancer effects. In particular, the potential cytotoxic activity may be attributed to secondary metabolites within the extract, such as alkaloids, flavonoids, and saponins, which are widely reported to exert cytotoxic effects on breast cancer cells [75-77]. Additional assays, including proliferation inhibition and apoptosis induction, are warranted to fully characterize the extract's therapeutic potential against breast cancer cells.
4. LIMITATIONS OF THIS STUDY
This study has several limitations that should be acknowledged. The in silico analyses were performed under simplified conditions that may not fully represent the biological environment. The in vitro evaluation was conducted on a single breast cancer cell line, MCF-7, which may limit the generalizability of the cytotoxicity results. Future studies involving multiple cell lines, mechanistic assays, and in vivo validation would strengthen and expand these findings.
CONCLUSION
WEE1 is a key regulator of the cell cycle and a potential therapeutic target, especially for breast cancer therapy. The ethanol extract of Ampelocissus thyrsiflora was found to contain flavonoids, glycosides, and steroids, known for their anticancer potential. Although some compounds did not meet physicochemical criteria, molecular docking and dynamics simulations were performed to explore their interactions with the WEE1 receptor. Of the 39 compounds identified, three demonstrated strong and stable interactions with WEE1, supported by favorable binding affinities and low MM-PBSA energy values. These compounds formed key hydrogen bonds with GLU 377 and CYS 379, and hydrophobic interactions with ALA 326 and CYS 379, residues critical for WEE1 inhibition. To support these findings, an in vitro MTT assay in MCF-7 cells confirmed a dose-dependent cytotoxic effect. Nonetheless, the combined in silico and in vitro results suggest that A. thyrsiflora metabolites hold promise as early-stage WEE1 inhibitors and merit further investigation.
AUTHORS’ CONTRIBUTIONS
The authors confirm contribution to the paper as follows: H.S.W.: Study conception and design; N.: Data collection; S.Y.: Analysis and interpretation of results; G.T.: Visualization; P.B.N.: Investigation; C.C.J.: Draft manuscript. All authors reviewed the results and approved the final version of the manuscript.
LIST OF ABBREVIATIONS
| GCO | = Global Cancer Observatory |
| CDK1 | = Cyclin Dependent-Kinase 1 |
| DNA | = Deoxyribonucleic Acid |
| BSLT | = Brine Shrimp Lethality Test |
| SIRT1 | = Sirtuin1 |
| AMPK | = AMP-Activated Protein Kinase |
| MTT | = 3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide |
| SMILES | = Simplified Molecular Input Line Entry System |
| MD | = Molecular Dynamics |
| RMSD | = Root Mean Square Deviation |
| RMSF | = Root Mean Square Fluctuation |
| SASA | = Solvent Accessible Surface Area |
| MMPBSA | = Molecular Mechanics Poisson-Boltzmann Surface Area |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Ethical approval was obtained from the Animal Research Ethics Committee of the Faculty of Mathematics and Natural Sciences, Universitas Sumatera Utara, Indonesia (Approval No. 0950/KEPH-FMIPA/2024).
HUMAN AND ANIMAL RIGHTS
All human research procedures followed were in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national), and with the Helsinki Declaration of 1975, as revised in 2013.
AVAILABILITY OF DATA AND MATERIALS
The data and supportive information are available within the article.
FUNDING
The authors would like to thank the TALENTA Research Program, Universitas Sumatera Utara, for funding this research through research contract No. 18589/UN5.1.R/PPM/2024, dated 30 May 2024.
ACKNOWLEDGEMENTS
The authors would like to thank the Faculty of Pharmacy, Universitas Sumatera, and the Faculty of Medicine, Public Health, and Nursing, Universitas Gadjah Mada, for their assistance and support, and for facilitating the study.


